Combining -Omics to Unravel the Impact of Copper Nutrition on Alfalfa (Medicago sativa) Stem Metabolism
- PMID: 26865661
- PMCID: PMC4771972
- DOI: 10.1093/pcp/pcw001
Combining -Omics to Unravel the Impact of Copper Nutrition on Alfalfa (Medicago sativa) Stem Metabolism
Abstract
Copper can be found in the environment at concentrations ranging from a shortage up to the threshold of toxicity for plants, with optimal growth conditions situated in between. The plant stem plays a central role in transferring and distributing minerals, water and other solutes throughout the plant. In this study, alfalfa is exposed to different levels of copper availability, from deficiency to slight excess, and the impact on the metabolism of the stem is assessed by a non-targeted proteomics study and by the expression analysis of key genes controlling plant stem development. Under copper deficiency, the plant stem accumulates specific copper chaperones, the expression of genes involved in stem development is decreased and the concentrations of zinc and molybdenum are increased in comparison with the optimum copper level. At the optimal copper level, the expression of cell wall-related genes increases and proteins playing a role in cell wall deposition and in methionine metabolism accumulate, whereas copper excess imposes a reduction in the concentration of iron in the stem and a reduced abundance of ferritins. Secondary ion mass spectrometry (SIMS) analysis suggests a role for the apoplasm as a copper storage site in the case of copper toxicity.
Keywords: ATX1; Cellulose synthase; Copper deficiency; Copper excess; Ionomics; Stem.
© The Author 2016. Published by Oxford University Press on behalf of Japanese Society of Plant Physiologists.
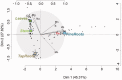
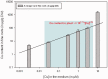
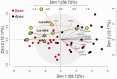
Figures


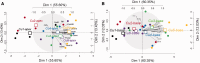



Similar articles
-
Long-term cadmium exposure influences the abundance of proteins that impact the cell wall structure in Medicago sativa stems.Plant Biol (Stuttg). 2018 Nov;20(6):1023-1035. doi: 10.1111/plb.12865. Epub 2018 Jul 24. Plant Biol (Stuttg). 2018. PMID: 29908008 Free PMC article.
-
Ups and downs in alfalfa: Proteomic and metabolic changes occurring in the growing stem.Plant Sci. 2015 Sep;238:13-25. doi: 10.1016/j.plantsci.2015.05.014. Epub 2015 May 22. Plant Sci. 2015. PMID: 26259170
-
Changes in the Proteome of Medicago sativa Leaves in Response to Long-Term Cadmium Exposure Using a Cell-Wall Targeted Approach.Int J Mol Sci. 2018 Aug 24;19(9):2498. doi: 10.3390/ijms19092498. Int J Mol Sci. 2018. PMID: 30149497 Free PMC article.
-
Effects of nitrogen source and water availability on stem carbohydrates and cellulosic bioethanol traits of alfalfa plants.Plant Sci. 2012 Aug;191-192:16-23. doi: 10.1016/j.plantsci.2012.04.007. Epub 2012 Apr 24. Plant Sci. 2012. PMID: 22682561
-
Transgene silencing of sucrose synthase in alfalfa (Medicago sativa L.) stem vascular tissue suggests a role for invertase in cell wall cellulose synthesis.BMC Plant Biol. 2015 Dec 1;15:283. doi: 10.1186/s12870-015-0649-4. BMC Plant Biol. 2015. PMID: 26627884 Free PMC article.
Cited by
-
Characterization of zinc uptake and translocation visualized with positron-emitting 65Zn tracer and analysis of transport-related gene expression in two Lotus japonicus accessions.Ann Bot. 2022 Dec 16;130(6):799-810. doi: 10.1093/aob/mcac101. Ann Bot. 2022. PMID: 35948001 Free PMC article.
-
Does long-term cadmium exposure influence the composition of pectic polysaccharides in the cell wall of Medicago sativa stems?BMC Plant Biol. 2019 Jun 21;19(1):271. doi: 10.1186/s12870-019-1859-y. BMC Plant Biol. 2019. PMID: 31226937 Free PMC article.
-
Biotechnological Perspectives of Omics and Genetic Engineering Methods in Alfalfa.Front Plant Sci. 2020 May 21;11:592. doi: 10.3389/fpls.2020.00592. eCollection 2020. Front Plant Sci. 2020. PMID: 32508859 Free PMC article. Review.
-
Long-Term Cd Exposure Alters the Metabolite Profile in Stem Tissue of Medicago sativa.Cells. 2020 Dec 17;9(12):2707. doi: 10.3390/cells9122707. Cells. 2020. PMID: 33348837 Free PMC article.
-
The Dynamics of the Cell Wall Proteome of Developing Alfalfa Stems.Biology (Basel). 2019 Aug 19;8(3):60. doi: 10.3390/biology8030060. Biology (Basel). 2019. PMID: 31430995 Free PMC article.
References
-
- Bouazizi H., Jouili H., Geitmann A., El Ferjani E. (2011) Cell wall accumulation of Cu ions and modulation of lignifying enzymes in primary leaves of bean seedlings exposed to excess copper. Biol Trace Elem. Res. 139: 97–107. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

