Novel KCNJ10 Gene Variations Compromise Function of Inwardly Rectifying Potassium Channel 4.1
- PMID: 26867573
- PMCID: PMC4817196
- DOI: 10.1074/jbc.M115.679910
Novel KCNJ10 Gene Variations Compromise Function of Inwardly Rectifying Potassium Channel 4.1
Abstract
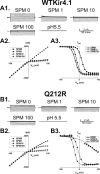
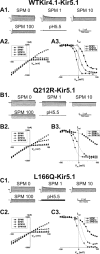
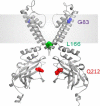
TheKCNJ10gene encoding Kir4.1 contains numerous SNPs whose molecular effects remain unknown. We investigated the functional consequences of uncharacterized SNPs (Q212R, L166Q, and G83V) on homomeric (Kir4.1) and heteromeric (Kir4.1-Kir5.1) channel function. We compared these with previously characterized EAST/SeSAME mutants (G77R and A167V) in kidney-derived tsA201 cells and in glial cell-derived C6 glioma cells. The membrane potentials of tsA201 cells expressing G77R and G83V were significantly depolarized as compared with WTKir4.1, whereas cells expressing Q212R, L166Q, and A167V were less affected. Furthermore, macroscopic currents from cells expressing WTKir4.1 and Q212R channels did not differ, whereas currents from cells expressing L166Q, G83V, G77R, and A167V were reduced. Unexpectedly, L166Q current responses were rescued when co-expressed with Kir5.1. In addition, we observed notable differences in channel activity between C6 glioma cells and tsA201 cells expressing L166Q and A167V, suggesting that there are underlying differences between cell lines in terms of Kir4.1 protein synthesis, stability, or expression at the surface. Finally, we determined spermine (SPM) sensitivity of these uncharacterized SNPs and found that Q212R-containing channels displayed reduced block by 1 μmSPM. At 100 μmSPM, the block was equal to or greater than WT, suggesting that the greater driving force of SPM allowed achievement of steady state. In contrast, L166Q-Kir5.1 channels achieved a higher block than WT, suggesting a more stable interaction of SPM in the deep pore cavity. Overall, our data suggest that G83V, L166Q, and Q212R residues play a pivotal role in controlling Kir4.1 channel function.
Keywords: KCNJ10; Kir4.1; epilepsy; kinetics; molecular cell biology; patch clamp; potassium channel; potassium transport; single nucleotide polymorphism (SNP); spermine.
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures





References
-
- Kucheryavykh Y. V., Kucheryavykh L. Y., Nichols C. G., Maldonado H. M., Baksi K., Reichenbach A., Skatchkov S. N., and Eaton M. J. (2007) Downregulation of Kir4.1 inward rectifying potassium channel subunits by RNAi impairs potassium transfer and glutamate uptake by cultured cortical astrocytes. Glia 55, 274–281 - PubMed
-
- Nichols C. G., and Lopatin A. N. (1997) Inward rectifier potassium channels. Annu. Rev. Physiol. 59, 171–191 - PubMed
-
- Bichet D., Haass F. A., and Jan L. Y. (2003) Merging functional studies with structures of inward-rectifier K+ channels. Nat. Rev. Neurosci. 4, 957–967 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

