Contributions of immunoaffinity chromatography to deep proteome profiling of human biofluids
- PMID: 26868616
- PMCID: PMC4862894
- DOI: 10.1016/j.jchromb.2016.01.015
Contributions of immunoaffinity chromatography to deep proteome profiling of human biofluids
Abstract
Human biofluids, especially blood plasma or serum, hold great potential as the sources of candidate biomarkers for various diseases; however, the enormous dynamic range of protein concentrations in biofluids represents a significant analytical challenge for detecting promising low-abundance proteins. Over the last decade, various immunoaffinity chromatographic methods have been developed and routinely applied for separating low-abundance proteins from the high- and moderate-abundance proteins, thus enabling much more effective detection of low-abundance proteins. Herein, we review the advances of immunoaffinity separation methods and their contributions to the proteomic applications in human biofluids. The limitations and future perspectives of immunoaffinity separation methods are also discussed.
Keywords: Affinity proteomics; Biofluid; Biomarker discovery; Immunoaffinity chromatography; Immunodepletion; Immunoenrichment; Plasma/Serum; Proteomics.
Copyright © 2016 Elsevier B.V. All rights reserved.
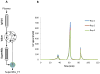
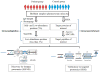
Figures
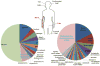

Similar articles
-
Immunoaffinity Depletion of High-Abundance Proteins from Serum/Plasma for Proteomic Analysis.Methods Mol Biol. 2025;2884:1-12. doi: 10.1007/978-1-0716-4298-6_1. Methods Mol Biol. 2025. PMID: 39715993
-
IgY14 and SuperMix immunoaffinity separations coupled with liquid chromatography-mass spectrometry for human plasma proteomics biomarker discovery.Methods. 2012 Feb;56(2):246-53. doi: 10.1016/j.ymeth.2011.09.001. Epub 2011 Sep 10. Methods. 2012. PMID: 21925605 Free PMC article. Review.
-
Enhanced Detection of Low-Abundance Human Plasma Proteins by Integrating Polyethylene Glycol Fractionation and Immunoaffinity Depletion.PLoS One. 2016 Nov 10;11(11):e0166306. doi: 10.1371/journal.pone.0166306. eCollection 2016. PLoS One. 2016. PMID: 27832179 Free PMC article.
-
Evaluation of multiprotein immunoaffinity subtraction for plasma proteomics and candidate biomarker discovery using mass spectrometry.Mol Cell Proteomics. 2006 Nov;5(11):2167-74. doi: 10.1074/mcp.T600039-MCP200. Epub 2006 Jul 19. Mol Cell Proteomics. 2006. PMID: 16854842 Free PMC article.
-
Emerging Affinity-Based Proteomic Technologies for Large-Scale Plasma Profiling in Cardiovascular Disease.Circulation. 2017 Apr 25;135(17):1651-1664. doi: 10.1161/CIRCULATIONAHA.116.025446. Circulation. 2017. PMID: 28438806 Free PMC article. Review.
Cited by
-
Proteomic Discovery and Validation of Novel Fluid Biomarkers for Improved Patient Selection and Prediction of Clinical Outcomes in Alzheimer's Disease Patient Cohorts.Proteomes. 2022 Aug 1;10(3):26. doi: 10.3390/proteomes10030026. Proteomes. 2022. PMID: 35997438 Free PMC article. Review.
-
Elucidating regulatory processes of intense physical activity by multi-omics analysis.Mil Med Res. 2023 Oct 18;10(1):48. doi: 10.1186/s40779-023-00477-5. Mil Med Res. 2023. PMID: 37853489 Free PMC article.
-
Comprehensive Analysis of RNA Expression Correlations between Biofluids and Human Tissues.Genes (Basel). 2021 Jun 18;12(6):935. doi: 10.3390/genes12060935. Genes (Basel). 2021. PMID: 34207420 Free PMC article.
-
Affinity depletion versus relative protein enrichment: a side-by-side comparison of two major strategies for increasing human cerebrospinal fluid proteome coverage.Clin Proteomics. 2019 Feb 26;16:9. doi: 10.1186/s12014-019-9229-1. eCollection 2019. Clin Proteomics. 2019. PMID: 30890900 Free PMC article.
-
Mass Spectrometry-Based Plasma Proteomics: Considerations from Sample Collection to Achieving Translational Data.J Proteome Res. 2019 Dec 6;18(12):4085-4097. doi: 10.1021/acs.jproteome.9b00503. Epub 2019 Oct 11. J Proteome Res. 2019. PMID: 31573204 Free PMC article. Review.
References
-
- Lathrop JT, Anderson NL, Anderson NG, Hammond DJ. Therapeutic potential of the plasma proteome. Curr Opin Mol Ther. 2003;5:250–257. - PubMed
-
- Anderson NL. The clinical plasma proteome: a survey of clinical assays for proteins in plasma and serum. Clin Chem. 2010;56:177–185. - PubMed
-
- Anderson NL, Anderson NG. The human plasma proteome: history, character, and diagnostic prospects. Mol Cell Proteomics. 2002;1:845–867. - PubMed
-
- Jacobs JM, Adkins JN, Qian WJ, Liu T, Shen Y, Camp DG, 2nd, Smith RD. Utilizing human blood plasma for proteomic biomarker discovery. J Proteome Res. 2005;4:1073–1085. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

