Widespread Expansion of Protein Interaction Capabilities by Alternative Splicing
- PMID: 26871637
- PMCID: PMC4882190
- DOI: 10.1016/j.cell.2016.01.029
Widespread Expansion of Protein Interaction Capabilities by Alternative Splicing
Abstract
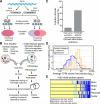
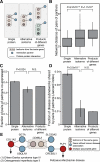
While alternative splicing is known to diversify the functional characteristics of some genes, the extent to which protein isoforms globally contribute to functional complexity on a proteomic scale remains unknown. To address this systematically, we cloned full-length open reading frames of alternatively spliced transcripts for a large number of human genes and used protein-protein interaction profiling to functionally compare hundreds of protein isoform pairs. The majority of isoform pairs share less than 50% of their interactions. In the global context of interactome network maps, alternative isoforms tend to behave like distinct proteins rather than minor variants of each other. Interaction partners specific to alternative isoforms tend to be expressed in a highly tissue-specific manner and belong to distinct functional modules. Our strategy, applicable to other functional characteristics, reveals a widespread expansion of protein interaction capabilities through alternative splicing and suggests that many alternative "isoforms" are functionally divergent (i.e., "functional alloforms").
Copyright © 2016 Elsevier Inc. All rights reserved.
Figures


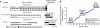
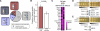
Comment in
-
SYSTEMS BIOLOGY. Protein isoforms: more than meets the eye.Nat Methods. 2016 Apr;13(4):291. doi: 10.1038/nmeth.3828. Nat Methods. 2016. PMID: 27482571 No abstract available.
References
-
- Barbosa-Morais NL, Irimia M, Pan Q, Xiong HY, Gueroussov S, Lee LJ, Slobodeniuc V, Kutter C, Watt S, Colak R, et al. The evolutionary landscape of alternative splicing in vertebrate species. Science. 2012;338:1587–1593. - PubMed
-
- Blencowe BJ. Alternative splicing: new insights from global analyses. Cell. 2006;126:37–47. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P50HG004233/HG/NHGRI NIH HHS/United States
- R01 HG001715/HG/NHGRI NIH HHS/United States
- R01 HD065288/HD/NICHD NIH HHS/United States
- T32CA009361/CA/NCI NIH HHS/United States
- R01 GM105431/GM/NIGMS NIH HHS/United States
- R01 MH091350/MH/NIMH NIH HHS/United States
- R33CA132073/CA/NCI NIH HHS/United States
- U01 HG001715/HG/NHGRI NIH HHS/United States
- P50 HG004233/HG/NHGRI NIH HHS/United States
- R21 MH104766/MH/NIMH NIH HHS/United States
- R33 CA132073/CA/NCI NIH HHS/United States
- R21MH104766/MH/NIMH NIH HHS/United States
- R01 MH105524/MH/NIMH NIH HHS/United States
- R01MH091350/MH/NIMH NIH HHS/United States
- U41 HG001715/HG/NHGRI NIH HHS/United States
- R01HD065288/HD/NICHD NIH HHS/United States
- R01MH105524/MH/NIMH NIH HHS/United States
- U01HG001715/HG/NHGRI NIH HHS/United States
- T32 CA009361/CA/NCI NIH HHS/United States
- R01GM105431/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

