Transcription-associated processes cause DNA double-strand breaks and translocations in neural stem/progenitor cells
- PMID: 26873106
- PMCID: PMC4776469
- DOI: 10.1073/pnas.1525564113
Transcription-associated processes cause DNA double-strand breaks and translocations in neural stem/progenitor cells
Abstract
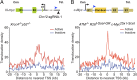
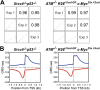
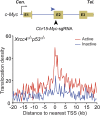
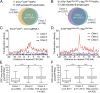
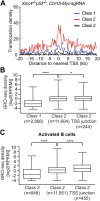
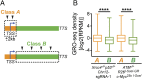
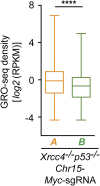
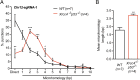
High-throughput, genome-wide translocation sequencing (HTGTS) studies of activated B cells have revealed that DNA double-strand breaks (DSBs) capable of translocating to defined bait DSBs are enriched around the transcription start sites (TSSs) of active genes. We used the HTGTS approach to investigate whether a similar phenomenon occurs in primary neural stem/progenitor cells (NSPCs). We report that breakpoint junctions indeed are enriched around TSSs that were determined to be active by global run-on sequencing analyses of NSPCs. Comparative analyses of transcription profiles in NSPCs and B cells revealed that the great majority of TSS-proximal junctions occurred in genes commonly expressed in both cell types, possibly because this common set has higher transcription levels on average than genes transcribed in only one or the other cell type. In the latter context, among all actively transcribed genes containing translocation junctions in NSPCs, those with junctions located within 2 kb of the TSS show a significantly higher transcription rate on average than genes with junctions in the gene body located at distances greater than 2 kb from the TSS. Finally, analysis of repair junction signatures of TSS-associated translocations in wild-type versus classical nonhomologous end-joining (C-NHEJ)-deficient NSPCs reveals that both C-NHEJ and alternative end-joining pathways can generate translocations by joining TSS-proximal DSBs to DSBs on other chromosomes. Our studies show that the generation of transcription-associated DSBs is conserved across divergent cell types.
Keywords: alternative end-joining; neural stem cells; nonhomologous end-joining; transcription; translocation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
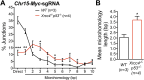
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

