Subtype-Specific Genes that Characterize Subpopulations of Callosal Projection Neurons in Mouse Identify Molecularly Homologous Populations in Macaque Cortex
- PMID: 26874185
- PMCID: PMC6317451
- DOI: 10.1093/cercor/bhw023
Subtype-Specific Genes that Characterize Subpopulations of Callosal Projection Neurons in Mouse Identify Molecularly Homologous Populations in Macaque Cortex
Abstract
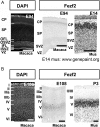
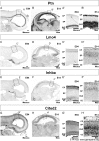
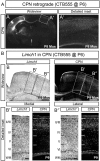
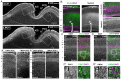
Callosal projection neurons (CPN) interconnect the neocortical hemispheres via the corpus callosum and are implicated in associative integration of multimodal information. CPN have undergone differential evolutionary elaboration, leading to increased diversity of cortical neurons-and more extensive and varied connections in neocortical gray and white matter-in primates compared with rodents. In mouse, distinct sets of genes are enriched in discrete subpopulations of CPN, indicating the molecular diversity of rodent CPN. Elements of rodent CPN functional and organizational diversity might thus be present in the further elaborated primate cortex. We address the hypothesis that genes controlling mouse CPN subtype diversity might reflect molecular patterns shared among mammals that arose prior to the divergence of rodents and primates. We find that, while early expression of the examined CPN-enriched genes, and postmigratory expression of these CPN-enriched genes in deep layers are highly conserved (e.g., Ptn, Nnmt, Cited2, Dkk3), in contrast, the examined genes expressed by superficial layer CPN show more variable levels of conservation (e.g., EphA3, Chn2). These results suggest that there has been evolutionarily differential retraction and elaboration of superficial layer CPN subpopulations between mouse and macaque, with independent derivation of novel populations in primates. Together, these data inform future studies regarding CPN subpopulations that are unique to primates and rodents, and indicate putative evolutionary relationships.
Keywords: corpus callosum; development; evolution; primate; rodent.
© The Author 2016. Published by Oxford University Press. All rights reserved. For Permissions, please e-mail: journals.permissions@oup.com.
Figures



Similar articles
-
New Molecular Players in the Development of Callosal Projections.Cells. 2020 Dec 26;10(1):29. doi: 10.3390/cells10010029. Cells. 2020. PMID: 33375263 Free PMC article. Review.
-
Caveolin1 Identifies a Specific Subpopulation of Cerebral Cortex Callosal Projection Neurons (CPN) Including Dual Projecting Cortical Callosal/Frontal Projection Neurons (CPN/FPN).eNeuro. 2018 Jan 18;5(1):ENEURO.0234-17.2017. doi: 10.1523/ENEURO.0234-17.2017. eCollection 2018 Jan-Feb. eNeuro. 2018. PMID: 29379878 Free PMC article.
-
Cited2 Regulates Neocortical Layer II/III Generation and Somatosensory Callosal Projection Neuron Development and Connectivity.J Neurosci. 2016 Jun 15;36(24):6403-19. doi: 10.1523/JNEUROSCI.4067-15.2016. J Neurosci. 2016. PMID: 27307230 Free PMC article.
-
Novel subtype-specific genes identify distinct subpopulations of callosal projection neurons.J Neurosci. 2009 Sep 30;29(39):12343-54. doi: 10.1523/JNEUROSCI.6108-08.2009. J Neurosci. 2009. PMID: 19793993 Free PMC article.
-
Development, specification, and diversity of callosal projection neurons.Trends Neurosci. 2011 Jan;34(1):41-50. doi: 10.1016/j.tins.2010.10.002. Epub 2010 Dec 2. Trends Neurosci. 2011. PMID: 21129791 Free PMC article. Review.
Cited by
-
Transcription factor expression defines subclasses of developing projection neurons highly similar to single-cell RNA-seq subtypes.Proc Natl Acad Sci U S A. 2020 Oct 6;117(40):25074-25084. doi: 10.1073/pnas.2008013117. Epub 2020 Sep 18. Proc Natl Acad Sci U S A. 2020. PMID: 32948690 Free PMC article.
-
Synthetic modified Fezf2 mRNA (modRNA) with concurrent small molecule SIRT1 inhibition enhances refinement of cortical subcerebral/corticospinal neuron identity from mouse embryonic stem cells.PLoS One. 2021 Sep 2;16(9):e0254113. doi: 10.1371/journal.pone.0254113. eCollection 2021. PLoS One. 2021. PMID: 34473715 Free PMC article.
-
New Molecular Players in the Development of Callosal Projections.Cells. 2020 Dec 26;10(1):29. doi: 10.3390/cells10010029. Cells. 2020. PMID: 33375263 Free PMC article. Review.
-
Caveolin1 Identifies a Specific Subpopulation of Cerebral Cortex Callosal Projection Neurons (CPN) Including Dual Projecting Cortical Callosal/Frontal Projection Neurons (CPN/FPN).eNeuro. 2018 Jan 18;5(1):ENEURO.0234-17.2017. doi: 10.1523/ENEURO.0234-17.2017. eCollection 2018 Jan-Feb. eNeuro. 2018. PMID: 29379878 Free PMC article.
-
Cross-species single-cell transcriptomics reveals neuronal similarities and heterogeneity in amniote pallium.Zool Res. 2025 Jan 18;46(1):193-208. doi: 10.24272/j.issn.2095-8137.2024.102. Zool Res. 2025. PMID: 39846196 Free PMC article.
References
-
- Aboitiz F, Montiel J. 2003. One hundred million years of interhemispheric communication: the history of the corpus callosum. Braz J Med Biol Res. 36:409–420. - PubMed
-
- Aboitiz F, Morales D, Montiel J. 2003. The evolutionary origin of the mammalian isocortex: towards an integrated developmental and functional approach. Behav Brain Sci. 26:535–552; discussion 552–585. - PubMed
-
- Arlotta P, Molyneaux BJ, Chen J, Inoue J, Kominami R, Macklis JD. 2005. Neuronal subtype-specific genes that control corticospinal motor neuron development in vivo. Neuron. 45:207–221. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

