Inhibition of oncogenic BRAF activity by indole-3-carbinol disrupts microphthalmia-associated transcription factor expression and arrests melanoma cell proliferation
- PMID: 26878440
- PMCID: PMC4985490
- DOI: 10.1002/mc.22472
Inhibition of oncogenic BRAF activity by indole-3-carbinol disrupts microphthalmia-associated transcription factor expression and arrests melanoma cell proliferation
Abstract
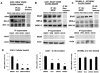
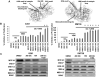
Indole-3-carbinol (I3C), an anti-cancer phytochemical derived from cruciferous vegetables, strongly inhibited proliferation and down-regulated protein levels of the melanocyte master regulator micropthalmia-associated transcription factor (MITF-M) in oncogenic BRAF-V600E expressing melanoma cells in culture as well as in vivo in tumor xenografted athymic nude mice. In contrast, wild type BRAF-expressing melanoma cells remained relatively insensitive to I3C anti-proliferative signaling. In BRAF-V600E-expressing melanoma cells, I3C treatment inhibited phosphorylation of MEK and ERK/MAPK, the down stream effectors of BRAF. The I3C anti-proliferative arrest was concomitant with the down-regulation of MITF-M transcripts and promoter activity, loss of endogenous BRN-2 binding to the MITF-M promoter, and was strongly attenuated by expression of exogenous MITF-M. Importantly, in vitro kinase assays using immunoprecipitated BRAF-V600E and wild type BRAF demonstrated that I3C selectively inhibited the enzymatic activity of the oncogenic BRAF-V600E but not of the wild type protein. In silico modeling predicted an I3C interaction site in the BRAF-V600E protomer distinct from where the clinically used BRAF-V600E inhibitor Vemurafenib binds to BRAF-V600E. Consistent with this prediction, combinations of I3C and Vemurafenib more potently inhibited melanoma cell proliferation and reduced MITF-M levels in BRAF-V600E expressing melanoma cells compared to the effects of each compound alone. Thus, our results demonstrate that oncogenic BRAF-V600E is a new cellular target of I3C that implicate this indolecarbinol compound as a potential candidate for novel single or combination therapies for melanoma. © 2016 Wiley Periodicals, Inc.
Keywords: BRN2; I3C; MITF-M; MITF-M promoter activity; Vemurafenib combinations; anti-proiferative signaling; human melanoma; indole-3-carbinol.
© 2016 Wiley Periodicals, Inc.
Figures



Similar articles
-
Cooperative antiproliferative signaling by aspirin and indole-3-carbinol targets microphthalmia-associated transcription factor gene expression and promoter activity in human melanoma cells.Cell Biol Toxicol. 2016 Apr;32(2):103-19. doi: 10.1007/s10565-016-9321-5. Epub 2016 Apr 7. Cell Biol Toxicol. 2016. PMID: 27055402 Free PMC article.
-
1-Benzyl-indole-3-carbinol is a highly potent new small molecule inhibitor of Wnt/β-catenin signaling in melanoma cells that coordinately inhibits cell proliferation and disrupts expression of microphthalmia-associated transcription factor isoform-M.Carcinogenesis. 2017 Dec 7;38(12):1207-1217. doi: 10.1093/carcin/bgx103. Carcinogenesis. 2017. PMID: 29028954 Free PMC article.
-
BRAFV600 inhibition alters the microRNA cargo in the vesicular secretome of malignant melanoma cells.Proc Natl Acad Sci U S A. 2017 Jul 18;114(29):E5930-E5939. doi: 10.1073/pnas.1705206114. Epub 2017 Jul 6. Proc Natl Acad Sci U S A. 2017. PMID: 28684402 Free PMC article.
-
Vemurafenib in patients with BRAF V600E mutation-positive advanced melanoma.Clin Ther. 2012 Jul;34(7):1474-86. doi: 10.1016/j.clinthera.2012.06.009. Epub 2012 Jun 27. Clin Ther. 2012. PMID: 22742884 Review.
-
Vemurafenib.Recent Results Cancer Res. 2014;201:215-25. doi: 10.1007/978-3-642-54490-3_13. Recent Results Cancer Res. 2014. PMID: 24756795 Review.
Cited by
-
Current results on the biological and pharmacological activities of Indole-3-carbinol.EXCLI J. 2018 Feb 6;17:181-185. doi: 10.17179/excli2017-1028. eCollection 2018. EXCLI J. 2018. PMID: 29743856 Free PMC article. No abstract available.
-
Cooperative antiproliferative signaling by aspirin and indole-3-carbinol targets microphthalmia-associated transcription factor gene expression and promoter activity in human melanoma cells.Cell Biol Toxicol. 2016 Apr;32(2):103-19. doi: 10.1007/s10565-016-9321-5. Epub 2016 Apr 7. Cell Biol Toxicol. 2016. PMID: 27055402 Free PMC article.
-
A PAX3/BRN2 rheostat controls the dynamics of BRAF mediated MITF regulation in MITFhigh /AXLlow melanoma.Pigment Cell Melanoma Res. 2019 Mar;32(2):280-291. doi: 10.1111/pcmr.12741. Epub 2018 Oct 19. Pigment Cell Melanoma Res. 2019. PMID: 30277012 Free PMC article.
-
Resistance to Gemcitabine in Pancreatic Ductal Adenocarcinoma: A Physiopathologic and Pharmacologic Review.Cancers (Basel). 2022 May 18;14(10):2486. doi: 10.3390/cancers14102486. Cancers (Basel). 2022. PMID: 35626089 Free PMC article. Review.
-
Recent Development in the Search for Epidermal Growth Factor Receptor (EGFR) Inhibitors based on the Indole Pharmacophore.Curr Top Med Chem. 2024;24(7):581-613. doi: 10.2174/0115680266264206231020111820. Curr Top Med Chem. 2024. PMID: 37909440 Review.
References
-
- Uzarska M, Czajkowski R, Schwartz RA, Bajek A, Zegarska B, Drewa T. Chemoprevention of skin melanoma: Facts and myths. Melanoma Res. 2013;23:426–433. - PubMed
-
- Garraway LA, Widlund HR, Rubin MA, et al. Integrative genomic analyses identify MITF as a lineage survival oncogene amplified in malignant melanoma. Nature. 2005;436:117–122. - PubMed
-
- Ugurel S, Houben R, Schrama D, et al. Microphthalmia-associated transcription factor gene amplification in metastatic melanoma is a prognostic marker for patient survival, but not a predictive marker for chemosensitivity and chemotherapy response. Clin Cancer Res. 2007;13:6344–6350. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

