Comparative and phylogenetic analysis of the mitochondrial genomes in basal hymenopterans
- PMID: 26879745
- PMCID: PMC4754708
- DOI: 10.1038/srep20972
Comparative and phylogenetic analysis of the mitochondrial genomes in basal hymenopterans
Abstract
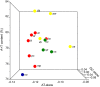
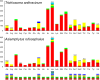
The Symphyta is traditionally accepted as a paraphyletic group located in a basal position of the order Hymenoptera. Herein, we conducted a comparative analysis of the mitochondrial genomes in the Symphyta by describing two newly sequenced ones, from Trichiosoma anthracinum, representing the first mitochondrial genome in family Cimbicidae, and Asiemphytus rufocephalus, from family Tenthredinidae. The sequenced lengths of these two mitochondrial genomes were 15,392 and 14,864 bp, respectively. Within the sequenced region, trnC and trnY were rearranged to the upstream of trnI-nad2 in T. anthracinum, while in A. rufocephalus all sequenced genes were arranged in the putative insect ancestral gene arrangement. Rearrangement of the tRNA genes is common in the Symphyta. The rearranged genes are mainly from trnL1 and two tRNA clusters of trnI-trnQ-trnM and trnW-trnC-trnY. The mitochondrial genomes of Symphyta show a biased usage of A and T rather than G and C. Protein-coding genes in Symphyta species show a lower evolutionary rate than those of Apocrita. The Ka/Ks ratios were all less than 1, indicating purifying selection of Symphyta species. Phylogenetic analyses supported the paraphyly and basal position of Symphyta in Hymenoptera. The well-supported phylogenetic relationship in the study is Tenthredinoidea + (Cephoidea + (Orussoidea + Apocrita)).
Figures




Similar articles
-
The mitochondrial genome of Tenthredo tienmushana (Takeuchi) and a related phylogenetic analysis of the sawflies (Insecta: Hymenoptera).Mitochondrial DNA A DNA Mapp Seq Anal. 2016 Jul;27(4):2860-1. doi: 10.3109/19401736.2015.1053129. Epub 2015 Jul 2. Mitochondrial DNA A DNA Mapp Seq Anal. 2016. PMID: 26134345
-
Nearly complete mitogenome of hairy sawfly, Corynis lateralis (Brullé, 1832) (Hymenoptera: Cimbicidae): rearrangements in the IQM and ARNS1EF gene clusters.Genetica. 2017 Oct;145(4-5):341-350. doi: 10.1007/s10709-017-9969-7. Epub 2017 May 31. Genetica. 2017. PMID: 28567603
-
The first two mitochondrial genomes of wood wasps (Hymenoptera: Symphyta): Novel gene rearrangements and higher-level phylogeny of the basal hymenopterans.Int J Biol Macromol. 2019 Feb 15;123:1189-1196. doi: 10.1016/j.ijbiomac.2018.11.017. Epub 2018 Nov 5. Int J Biol Macromol. 2019. PMID: 30408451
-
Next-Generation Sequencing of Two Mitochondrial Genomes from Family Pompilidae (Hymenoptera: Vespoidea) Reveal Novel Patterns of Gene Arrangement.Int J Mol Sci. 2016 Oct 11;17(10):1641. doi: 10.3390/ijms17101641. Int J Mol Sci. 2016. PMID: 27727175 Free PMC article.
-
Accelerated evolution of mitochondrial but not nuclear genomes of Hymenoptera: new evidence from crabronid wasps.PLoS One. 2012;7(3):e32826. doi: 10.1371/journal.pone.0032826. Epub 2012 Mar 6. PLoS One. 2012. PMID: 22412929 Free PMC article.
Cited by
-
The First Two Complete Mitochondrial Genomes for the Subfamily Meligethinae (Coleoptera: Nitidulidae) and Implications for the Higher Phylogeny of Nitidulidae.Insects. 2024 Jan 12;15(1):57. doi: 10.3390/insects15010057. Insects. 2024. PMID: 38249063 Free PMC article.
-
Evolutionary timescale of chalcidoid wasps inferred from over one hundred mitochondrial genomes.Zool Res. 2023 May 18;44(3):467-482. doi: 10.24272/j.issn.2095-8137.2022.379. Zool Res. 2023. PMID: 36994537 Free PMC article.
-
Comparative Mitogenomic Analysis of Two Cuckoo Bees (Apoidea: Anthophila: Megachilidae) with Phylogenetic Implications.Insects. 2021 Jan 5;12(1):29. doi: 10.3390/insects12010029. Insects. 2021. PMID: 33466344 Free PMC article.
-
Discriminating lineages of Batrachochytrium dendrobatidis using quantitative PCR.Mol Ecol Resour. 2021 Jul;21(5):1452-1459. doi: 10.1111/1755-0998.13299. Epub 2021 Feb 12. Mol Ecol Resour. 2021. PMID: 33232563 Free PMC article.
-
Phylogenomic Analyses of the Tenthredinoidea Support the Familial Rank of Athaliidae (Insecta, Tenthredinoidea).Insects. 2022 Sep 21;13(10):858. doi: 10.3390/insects13100858. Insects. 2022. PMID: 36292806 Free PMC article.
References
-
- Simon C. et al. Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann. Entomol. Soc. Am. 87, 651–701 (1994).
-
- Cameron S. L. Insect mitochondrial genomics: implications for evolution and phylogeny. Annu. Rev. Entomol. 59, 95–117 (2014). - PubMed
-
- Curole J. P. & Kocher T. D. Mitogenomics: digging deeper with complete mitochondrial genomes. Trends. Ecol. Evol. 14, 394–398 (1999). - PubMed
-
- Lin C. P. & Danforth B. N. How do insect nuclear and mitochondrial gene substitution patterns differ? Insights from Bayesian analyses of combined datasets. Mol. Phylogen. Evol. 30, 686–702 (2004). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

