Conserved rates and patterns of transcription errors across bacterial growth states and lifestyles
- PMID: 26884158
- PMCID: PMC4812759
- DOI: 10.1073/pnas.1525329113
Conserved rates and patterns of transcription errors across bacterial growth states and lifestyles
Erratum in
-
Correction for Traverse and Ochman, Conserved rates and patterns of transcription errors across bacterial growth states and lifestyles.Proc Natl Acad Sci U S A. 2016 Jul 19;113(29):E4257-8. doi: 10.1073/pnas.1609677113. Epub 2016 Jul 11. Proc Natl Acad Sci U S A. 2016. PMID: 27402746 Free PMC article. No abstract available.
Abstract
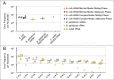
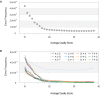
Errors that occur during transcription have received much less attention than the mutations that occur in DNA because transcription errors are not heritable and usually result in a very limited number of altered proteins. However, transcription error rates are typically several orders of magnitude higher than the mutation rate. Also, individual transcripts can be translated multiple times, so a single error can have substantial effects on the pool of proteins. Transcription errors can also contribute to cellular noise, thereby influencing cell survival under stressful conditions, such as starvation or antibiotic stress. Implementing a method that captures transcription errors genome-wide, we measured the rates and spectra of transcription errors in Escherichia coli and in endosymbionts for which mutation and/or substitution rates are greatly elevated over those of E. coli Under all tested conditions, across all species, and even for different categories of RNA sequences (mRNA and rRNAs), there were no significant differences in rates of transcription errors, which ranged from 2.3 × 10(-5) per nucleotide in mRNA of the endosymbiont Buchnera aphidicola to 5.2 × 10(-5) per nucleotide in rRNA of the endosymbiont Carsonella ruddii The similarity of transcription error rates in these bacterial endosymbionts to that in E. coli (4.63 × 10(-5) per nucleotide) is all the more surprising given that genomic erosion has resulted in the loss of transcription fidelity factors in both Buchnera and Carsonella.
Keywords: RNA polymerase fidelity; base substitutions; transcription errors.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
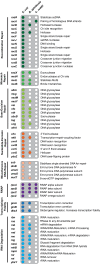
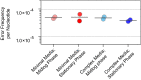
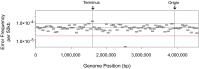
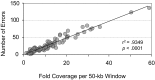


Comment in
-
Transcriptional errors and the drift barrier.Proc Natl Acad Sci U S A. 2016 Mar 22;113(12):3136-8. doi: 10.1073/pnas.1601785113. Epub 2016 Mar 10. Proc Natl Acad Sci U S A. 2016. PMID: 26966235 Free PMC article. No abstract available.
References
-
- Springgate CF, Loeb LA. On the fidelity of transcription by Escherichia coli ribonucleic acid polymerase. J Mol Biol. 1975;97(4):577–591. - PubMed
-
- Rosenberger RF, Foskett G. An estimate of the frequency of in vivo transcriptional errors at a nonsense codon in Escherichia coli. Mol Gen Genet. 1981;183(3):561–563. - PubMed
-
- Rosenberger RF, Hilton J. The frequency of transcriptional and translational errors at nonsense codons in the lacZ gene of Escherichia coli. Mol Gen Genet. 1983;191(2):207–212. - PubMed
-
- O’Farrell PH. The suppression of defective translation by ppGpp and its role in the stringent response. Cell. 1978;14(3):545–557. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

