A continuum model of transcriptional bursting
- PMID: 26896676
- PMCID: PMC4850746
- DOI: 10.7554/eLife.13051
A continuum model of transcriptional bursting
Abstract
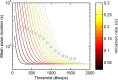
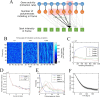
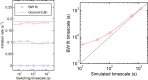
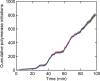
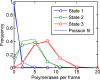
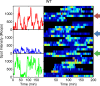
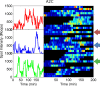
Transcription occurs in stochastic bursts. Early models based upon RNA hybridisation studies suggest bursting dynamics arise from alternating inactive and permissive states. Here we investigate bursting mechanism in live cells by quantitative imaging of actin gene transcription, combined with molecular genetics, stochastic simulation and probabilistic modelling. In contrast to early models, our data indicate a continuum of transcriptional states, with a slowly fluctuating initiation rate converting the gene between different levels of activity, interspersed with extended periods of inactivity. We place an upper limit of 40 s on the lifetime of fluctuations in elongation rate, with initiation rate variations persisting an order of magnitude longer. TATA mutations reduce the accessibility of high activity states, leaving the lifetime of on- and off-states unchanged. A continuum or spectrum of gene states potentially enables a wide dynamic range for cell responses to stimuli.
Keywords: chromosomes; computational biology; computational modelling; dictyostelium; genes; live cell imaging; single cell gene expression; stochastic gene expression; systems biology; transcription; transcriptional bursting.
Conflict of interest statement
The authors declare that no competing interests exist.
Figures
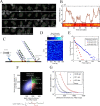
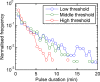
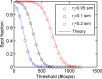
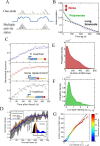


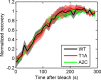
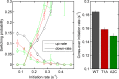



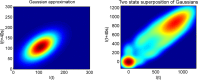
References
-
- Bothma JP, Garcia HG, Esposito E, Schlissel G, Gregor T, Levine M. Dynamic regulation of eve stripe 2 expression reveals transcriptional bursts in living Drosophila embryos. Proceedings of the National Academy of Sciences of the United States of America. 2014;111:10598–10603. doi: 10.1073/pnas.1410022111. - DOI - PMC - PubMed
-
- Burnham K, Anderson D. Model Selection and Multimodal Inference - a Practical Information-Theoretic Approach. 2nd. Springer; 2002.

