In silico selection of an aptamer to estrogen receptor alpha using computational docking employing estrogen response elements as aptamer-alike molecules
- PMID: 26899418
- PMCID: PMC4761961
- DOI: 10.1038/srep21285
In silico selection of an aptamer to estrogen receptor alpha using computational docking employing estrogen response elements as aptamer-alike molecules
Abstract
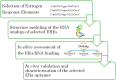
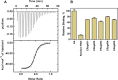
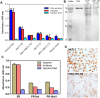
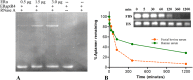
Aptamers, the chemical-antibody substitute to conventional antibodies, are primarily discovered through SELEX technology involving multi-round selections and enrichment. Circumventing conventional methodology, here we report an in silico selection of aptamers to estrogen receptor alpha (ERα) using RNA analogs of human estrogen response elements (EREs). The inverted repeat nature of ERE and the ability to form stable hairpins were used as criteria to obtain aptamer-alike sequences. Near-native RNA analogs of selected single stranded EREs were modelled and their likelihood to emerge as ERα aptamer was examined using AutoDock Vina, HADDOCK and PatchDock docking. These in silico predictions were validated by measuring the thermodynamic parameters of ERα -RNA interactions using isothermal titration calorimetry. Based on the in silico and in vitro results, we selected a candidate RNA (ERaptR4; 5'-GGGGUCAAGGUGACCCC-3') having a binding constant (Ka) of 1.02 ± 0.1 × 10(8) M(-1) as an ERα-aptamer. Target-specificity of the selected ERaptR4 aptamer was confirmed through cytochemistry and solid-phase immunoassays. Furthermore, stability analyses identified ERaptR4 resistant to serum and RNase A degradation in presence of ERα. Taken together, an efficient ERα-RNA aptamer is identified using a non-SELEX procedure of aptamer selection. The high-affinity and specificity can be utilized in detection of ERα in breast cancer and related diseases.
Figures

Similar articles
-
Aptamer-Assisted Detection of the Altered Expression of Estrogen Receptor Alpha in Human Breast Cancer.PLoS One. 2016 Apr 4;11(4):e0153001. doi: 10.1371/journal.pone.0153001. eCollection 2016. PLoS One. 2016. PMID: 27043307 Free PMC article.
-
In search of novel drug target sites on estrogen receptors using RNA aptamers.Nucleic Acid Ther. 2014 Jun;24(3):226-38. doi: 10.1089/nat.2013.0474. Epub 2014 Mar 3. Nucleic Acid Ther. 2014. PMID: 24588102 Free PMC article.
-
An improved SELEX technique for selection of DNA aptamers binding to M-type 11 of Streptococcus pyogenes.Methods. 2016 Mar 15;97:51-7. doi: 10.1016/j.ymeth.2015.12.005. Epub 2015 Dec 8. Methods. 2016. PMID: 26678795
-
In silico molecular docking in DNA aptamer development.Biochimie. 2021 Jan;180:54-67. doi: 10.1016/j.biochi.2020.10.005. Epub 2020 Oct 18. Biochimie. 2021. PMID: 33086095 Review.
-
In silico approaches to RNA aptamer design.Biochimie. 2018 Feb;145:8-14. doi: 10.1016/j.biochi.2017.10.005. Epub 2017 Oct 12. Biochimie. 2018. PMID: 29032056 Review.
Cited by
-
Potential Inherent Stimulation of the Innate Immune System by Nucleic Acid Aptamers and Possible Corrective Approaches.Pharmaceuticals (Basel). 2018 Jun 23;11(3):62. doi: 10.3390/ph11030062. Pharmaceuticals (Basel). 2018. PMID: 29937498 Free PMC article. Review.
-
Emerging cancer-specific therapeutic aptamers.Curr Opin Oncol. 2017 Sep;29(5):366-374. doi: 10.1097/CCO.0000000000000389. Curr Opin Oncol. 2017. PMID: 28692589 Free PMC article. Review.
-
Protein Network Studies on PCOS Biomarkers With S100A8, Druggability Assessment, and RNA Aptamer Designing to Control Its Cyst Migration Effect.Front Bioeng Biotechnol. 2020 May 13;8:328. doi: 10.3389/fbioe.2020.00328. eCollection 2020. Front Bioeng Biotechnol. 2020. PMID: 32478041 Free PMC article.
-
Distribution and Effects of Estrogen Receptors in Prostate Cancer: Associated Molecular Mechanisms.Front Endocrinol (Lausanne). 2022 Jan 11;12:811578. doi: 10.3389/fendo.2021.811578. eCollection 2021. Front Endocrinol (Lausanne). 2022. PMID: 35087479 Free PMC article. Review.
-
Predicting aptamer sequences that interact with target proteins using an aptamer-protein interaction classifier and a Monte Carlo tree search approach.PLoS One. 2021 Jun 25;16(6):e0253760. doi: 10.1371/journal.pone.0253760. eCollection 2021. PLoS One. 2021. PMID: 34170922 Free PMC article.
References
-
- Weigel M. T. & Dowsett M. Current and emerging biomarkers in breast cancer: prognosis and prediction. Endocr Relat Cancer 17, R245–262 (2010). - PubMed
-
- Ali S. & Coombes R. C. Estrogen receptor alpha in human breast cancer: Occurrence and significance. J Mammary Gland Biol Neoplasia 5, 271–281 (2000). - PubMed
-
- Lippman M. E. & Allegra J. C. Quantitative estrogen receptor analysis: The response to endocrine and cytotoxic chemotherapy in human breast cancer and the disease-free interval. Cancer 46, 2829–2834 (1980). - PubMed
-
- Fisher B. et al. Tamoxifen for prevention of breast cancer: report of the National Surgical Adjuvant Breast and Bowel Project P-1 Study. J Natl Cancer Inst 90, 1371–1388 (1998). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

