Knock-down of a RING finger gene confers cold tolerance
- PMID: 26901100
- PMCID: PMC4878256
- DOI: 10.1080/21655979.2015.1131368
Knock-down of a RING finger gene confers cold tolerance
Abstract
The plant-specific RING-domain finger proteins play important roles in plant development and stress responses. We recently identified and functionally characterized a stress-induced gene OsSRFP1 (Oryza sativa Stress-related RING Finger Protein 1) from rice. We showed evidences of the biotechnological potential of the suppression of OsSRFP1 expression in conferring cold tolerance. The increased cold tolerance of OsSRFP1 knock-down plants was associated with higher amounts of free proline and activities of antioxidant enzymes. In vitro ubiquitination assays showed that OsSRFP1 possessed E3 ubiquitin ligase activity. Some predicted interacting partners of OsSRFP1 might be the substrates for OsSRFP1-mediated protein degradation. Interestingly, OsSRFP1 had trans-activation activity, suggesting the dual roles of OsSRFP1 in post-translational and transcriptional regulations in stress responses.
Keywords: E3 ubiquitin ligase; cold stress; rice; transcription factor.
Figures




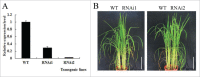
Erratum for
- doi: 10.1007/s11103-015-0294-1
Similar articles
-
Knock-down of stress inducible OsSRFP1 encoding an E3 ubiquitin ligase with transcriptional activation activity confers abiotic stress tolerance through enhancing antioxidant protection in rice.Plant Mol Biol. 2015 Mar;87(4-5):441-58. doi: 10.1007/s11103-015-0294-1. Epub 2015 Feb 11. Plant Mol Biol. 2015. PMID: 25667045
-
Oryza sativa heat-induced RING finger protein 1 (OsHIRP1) positively regulates plant response to heat stress.Plant Mol Biol. 2019 Apr;99(6):545-559. doi: 10.1007/s11103-019-00835-9. Epub 2019 Feb 7. Plant Mol Biol. 2019. PMID: 30730020
-
E3 ligase, the Oryza sativa salt-induced RING finger protein 4 (OsSIRP4), negatively regulates salt stress responses via degradation of the OsPEX11-1 protein.Plant Mol Biol. 2021 Feb;105(3):231-245. doi: 10.1007/s11103-020-01084-x. Epub 2020 Oct 20. Plant Mol Biol. 2021. PMID: 33079323
-
Research Progress on Plant RING-Finger Proteins.Genes (Basel). 2019 Nov 26;10(12):973. doi: 10.3390/genes10120973. Genes (Basel). 2019. PMID: 31779262 Free PMC article. Review.
-
Abiotic stress tolerance mediated by protein ubiquitination.J Exp Bot. 2012 Jan;63(2):599-616. doi: 10.1093/jxb/err310. Epub 2011 Oct 20. J Exp Bot. 2012. PMID: 22016431 Review.
Cited by
-
Stress-responsive gene RsICE1 from Raphanus sativus increases cold tolerance in rice.Protoplasma. 2017 Mar;254(2):945-956. doi: 10.1007/s00709-016-1004-9. Epub 2016 Jul 29. Protoplasma. 2017. PMID: 27473592
-
Integrated sRNA-seq and RNA-seq Analyses Reveal a microRNA Regulation Network Involved in Cold Response in Pisum sativum L.Genes (Basel). 2022 Jun 22;13(7):1119. doi: 10.3390/genes13071119. Genes (Basel). 2022. PMID: 35885902 Free PMC article.
-
Combined QTL mapping and RNA-Seq pro-filing reveal candidate genes related to low-temperature tolerance in maize.Mol Breed. 2022 Jun 20;42(6):33. doi: 10.1007/s11032-022-01297-6. eCollection 2022 Jun. Mol Breed. 2022. PMID: 37312966 Free PMC article.
-
Genome-Wide Association Analysis of Freezing Tolerance and Winter Hardiness in Winter Wheat of Nordic Origin.Plants (Basel). 2023 Nov 29;12(23):4014. doi: 10.3390/plants12234014. Plants (Basel). 2023. PMID: 38068649 Free PMC article.
-
Modulation of Abiotic Stress Responses in Rice by E3-Ubiquitin Ligases: A Promising Way to Develop Stress-Tolerant Crops.Front Plant Sci. 2021 Mar 23;12:640193. doi: 10.3389/fpls.2021.640193. eCollection 2021. Front Plant Sci. 2021. PMID: 33833769 Free PMC article. Review.
References
-
- Fang H, Meng Q, Xu J, Tang H, Tang S, Zhang H, Huang J. Knock-down of stress inducible OsSRFP1 encoding an E3 ubiquitin ligase with transcriptional activation activity confers abiotic stress tolerance through enhancing antioxidant protection in rice. Plant Mol Biol 2015; 87:441-58; PMID:25667045; http://dx.doi.org/10.1007/s11103-015-0294-1 - DOI - PubMed
-
- Huertas R, Olias R, Eljakaoui Z, Galvez FJ, Li J, De Morales PA, Belver A, Rodríguez-Rosales MP. Overexpression of SlSOS2 (SlCIPK24) confers salt tolerance to transgenic tomato. Plant, Cell Env 2012; 35:1467-82; PMID:22390672; http://dx.doi.org/10.1111/j.1365-3040.2012.02504.x - DOI - PubMed
-
- Belver A, Olias R, Huertas R, Rodriguez-Rosales MP. Involvement of SlSOS2 in tomato salt tolerance. Bioengineered 2012; 3:298-302; PMID:22825351; http://dx.doi.org/10.4161/bioe.20796 - DOI - PMC - PubMed
-
- Ma X, Zhu X, Li C, Song Y, Zhang W, Xia G, Wang M. Overexpression of wheat NF-YA10 gene regulates the salinity stress response in Arabidopsis thaliana. Plant Physiol Biochem 2015; 86:34-43; PMID:25461698; http://dx.doi.org/10.1016/j.plaphy.2014.11.011 - DOI - PubMed
-
- Ma X, Li C, Wang M. Wheat NF-YA10 functions independently in salinity and drought stress. Bioengineered 2015; 6:245-7; PMID:26083807; http://dx.doi.org/10.1080/21655979.2015.1054085 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
