Perspective: Adhesion Mediated Signal Transduction in Bacterial Pathogens
- PMID: 26901228
- PMCID: PMC4810144
- DOI: 10.3390/pathogens5010023
Perspective: Adhesion Mediated Signal Transduction in Bacterial Pathogens
Abstract
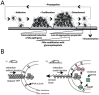
During the infection process, pathogenic bacteria undergo large-scale transcriptional changes to promote virulence and increase intrahost survival. While much of this reprogramming occurs in response to changes in chemical environment, such as nutrient availability and pH, there is increasing evidence that adhesion to host-tissue can also trigger signal transduction pathways resulting in differential gene expression. Determining the molecular mechanisms of adhesion-mediated signaling requires disentangling the contributions of chemical and mechanical stimuli. Here we highlight recent work demonstrating that surface attachment drives a transcriptional response in bacterial pathogens, including uropathogenic Escherichia coli (E. coli), and discuss the complexity of experimental design when dissecting the specific role of adhesion-mediated signaling during infection.
Keywords: adhesion; fimbriae; signal transduction; uropathogenic E. coli; virulence.
Figures


Similar articles
-
Altered Regulation of the Diguanylate Cyclase YaiC Reduces Production of Type 1 Fimbriae in a Pst Mutant of Uropathogenic Escherichia coli CFT073.J Bacteriol. 2017 Nov 14;199(24):e00168-17. doi: 10.1128/JB.00168-17. Print 2017 Dec 15. J Bacteriol. 2017. PMID: 28924030 Free PMC article.
-
Virulence factors of uropathogenic E. coli and their interaction with the host.Adv Microb Physiol. 2014;65:337-72. doi: 10.1016/bs.ampbs.2014.08.006. Epub 2014 Nov 4. Adv Microb Physiol. 2014. PMID: 25476769 Review.
-
Epithelial C5aR1 Signaling Enhances Uropathogenic Escherichia coli Adhesion to Human Renal Tubular Epithelial Cells.Front Immunol. 2018 May 1;9:949. doi: 10.3389/fimmu.2018.00949. eCollection 2018. Front Immunol. 2018. PMID: 29765378 Free PMC article.
-
Bacterial pathogenesis of plants: future challenges from a microbial perspective: Challenges in Bacterial Molecular Plant Pathology.Mol Plant Pathol. 2016 Oct;17(8):1298-313. doi: 10.1111/mpp.12427. Epub 2016 Aug 4. Mol Plant Pathol. 2016. PMID: 27170435 Free PMC article. Review.
-
Fimbriae-mediated host-pathogen cross-talk.Curr Opin Microbiol. 1998 Feb;1(1):75-81. doi: 10.1016/s1369-5274(98)80145-8. Curr Opin Microbiol. 1998. PMID: 10066469 Review.
Cited by
-
Against the tide: the role of bacterial adhesion in host colonization.Biochem Soc Trans. 2016 Dec 15;44(6):1571-1580. doi: 10.1042/BST20160186. Biochem Soc Trans. 2016. PMID: 27913666 Free PMC article. Review.
-
Arginine within a specific motif near the N-terminal of FimY is critical for the maximal production of type 1 fimbriae in Salmonella enterica serovar Typhimurium.Microbiologyopen. 2019 Sep;8(9):e00846. doi: 10.1002/mbo3.846. Epub 2019 Apr 16. Microbiologyopen. 2019. PMID: 30993839 Free PMC article.
-
Prediction of Broad-Spectrum Pathogen Attachment to Coating Materials for Biomedical Devices.ACS Appl Mater Interfaces. 2018 Jan 10;10(1):139-149. doi: 10.1021/acsami.7b14197. Epub 2018 Jan 2. ACS Appl Mater Interfaces. 2018. PMID: 29191009 Free PMC article.
-
Analysis of the contribution of MTP and the predicted Flp pilus genes to Mycobacterium tuberculosis pathogenesis.Microbiology (Reading). 2016 Oct;162(10):1784-1796. doi: 10.1099/mic.0.000368. Epub 2016 Aug 31. Microbiology (Reading). 2016. PMID: 27586540 Free PMC article.
-
Drosophila melanogaster establishes a species-specific mutualistic interaction with stable gut-colonizing bacteria.PLoS Biol. 2018 Jul 5;16(7):e2005710. doi: 10.1371/journal.pbio.2005710. eCollection 2018 Jul. PLoS Biol. 2018. PMID: 29975680 Free PMC article.
References
-
- Subashchandrabose S., Hazen T.H., Brumbaugh A.R., Himpsl S.D., Smith S.N., Ernst R.D., Rasko D.A., Mobley H.L. Host-specific induction of Escherichia coli fitness genes during human urinary tract infection. Proc. Natl. Acad. Sci. USA. 2014;111:18327–18332. doi: 10.1073/pnas.1415959112. - DOI - PMC - PubMed
-
- Mavromatis C.H., Bokil N.J., Totsika M., Kakkanat A., Schaale K., Cannistraci C.V., Ryu T., Beatson S.A., Ulett G.C., Schembri M.A., et al. The co-transcriptome of uropathogenic Escherichia coli-infected mouse macrophages reveals new insights into host-pathogen interactions. Cell. Microbiol. 2015;17:730–746. doi: 10.1111/cmi.12397. - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

