An Unusual Member of the Papain Superfamily: Mapping the Catalytic Cleft of the Marasmius oreades agglutinin (MOA) with a Caspase Inhibitor
- PMID: 26901797
- PMCID: PMC4764322
- DOI: 10.1371/journal.pone.0149407
An Unusual Member of the Papain Superfamily: Mapping the Catalytic Cleft of the Marasmius oreades agglutinin (MOA) with a Caspase Inhibitor
Abstract
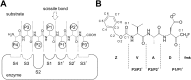
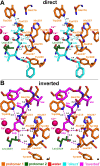
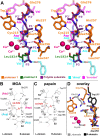
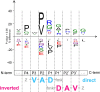
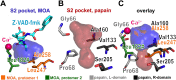
Papain-like cysteine proteases (PLCPs) constitute the largest group of thiol-based protein degrading enzymes and are characterized by a highly conserved fold. They are found in bacteria, viruses, plants and animals and involved in a number of physiological and pathological processes, parasitic infections and host defense, making them interesting targets for drug design. The Marasmius oreades agglutinin (MOA) is a blood group B-specific fungal chimerolectin with calcium-dependent proteolytic activity. The proteolytic domain of MOA presents a unique structural arrangement, yet mimicking the main structural elements in known PLCPs. Here we present the X-ray crystal structure of MOA in complex with Z-VAD-fmk, an irreversible caspase inhibitor known to cross-react with PLCPs. The structural data allow modeling of the substrate binding geometry and mapping of the fundamental enzyme-substrate interactions. The new information consolidates MOA as a new, yet strongly atypical member of the papain superfamily. The reported complex is the first published structure of a PLCP in complex with the well characterized caspase inhibitor Z-VAD-fmk.
Conflict of interest statement
Figures


Similar articles
-
Marasmius oreades agglutinin (MOA) is a chimerolectin with proteolytic activity.Biochem Biophys Res Commun. 2011 May 13;408(3):405-10. doi: 10.1016/j.bbrc.2011.04.031. Epub 2011 Apr 12. Biochem Biophys Res Commun. 2011. PMID: 21513701
-
Nematotoxicity of Marasmius oreades agglutinin (MOA) depends on glycolipid binding and cysteine protease activity.J Biol Chem. 2011 Sep 2;286(35):30337-30343. doi: 10.1074/jbc.M111.258202. Epub 2011 Jul 8. J Biol Chem. 2011. PMID: 21757752 Free PMC article.
-
Family of Papain-Like Fungal Chimerolectins with Distinct Ca2+-Dependent Activation Mechanism.Biochemistry. 2017 Sep 5;56(35):4689-4700. doi: 10.1021/acs.biochem.7b00317. Epub 2017 Aug 22. Biochemistry. 2017. PMID: 28665586
-
Papain-like cysteine proteases as hubs in plant immunity.New Phytol. 2016 Dec;212(4):902-907. doi: 10.1111/nph.14117. Epub 2016 Aug 4. New Phytol. 2016. PMID: 27488095 Review.
-
Role of Papain-Like Cysteine Proteases in Plant Development.Front Plant Sci. 2018 Dec 4;9:1717. doi: 10.3389/fpls.2018.01717. eCollection 2018. Front Plant Sci. 2018. PMID: 30564252 Free PMC article. Review.
Cited by
-
Crystal structure of MOA in complex with a peptide fragment: A protease caught in flagranti.Curr Res Struct Biol. 2020 Apr 22;2:56-67. doi: 10.1016/j.crstbi.2020.04.003. eCollection 2020. Curr Res Struct Biol. 2020. PMID: 34235469 Free PMC article.
-
The Crystal Structure of Bacillus thuringiensis Tpp80Aa1 and Its Interaction with Galactose-Containing Glycolipids.Toxins (Basel). 2022 Dec 8;14(12):863. doi: 10.3390/toxins14120863. Toxins (Basel). 2022. PMID: 36548760 Free PMC article.
-
Extending Janus lectins architecture: Characterization and application to protocells.Comput Struct Biotechnol J. 2022 Nov 7;20:6108-6119. doi: 10.1016/j.csbj.2022.11.005. eCollection 2022. Comput Struct Biotechnol J. 2022. PMID: 36420169 Free PMC article.
-
Marasmius oreades agglutinin enhances resistance of Arabidopsis against plant-parasitic nematodes and a herbivorous insect.BMC Plant Biol. 2021 Sep 1;21(1):402. doi: 10.1186/s12870-021-03186-0. BMC Plant Biol. 2021. PMID: 34470613 Free PMC article.
References
-
- Turk D, Gunčar G, Podobnik M, Turk B. Revised definition of substrate binding sites of papain-like cysteine proteases. Biol Chem. 1998;379(2):137–47. Epub 1998/04/02. . - PubMed
-
- Turk B, Turk V, Turk D. Structural and functional aspects of papain-like cysteine proteinases and their protein inhibitors. Biol Chem. 1997;378(3–4):141–50. Epub 1997/03/01. . - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

