In silico Analysis of HIV-1 Env-gp120 Reveals Structural Bases for Viral Adaptation in Growth-Restrictive Cells
- PMID: 26903989
- PMCID: PMC4746247
- DOI: 10.3389/fmicb.2016.00110
In silico Analysis of HIV-1 Env-gp120 Reveals Structural Bases for Viral Adaptation in Growth-Restrictive Cells
Abstract
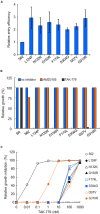
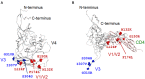
Variable V1/V2 and V3 loops on human immunodeficiency virus type 1 (HIV-1) envelope-gp120 core play key roles in modulating viral competence to recognize two infection receptors, CD4 and chemokine-receptors. However, molecular bases for the modulation largely remain unclear. To address these issues, we constructed structural models for a full-length gp120 in CD4-free and -bound states. The models showed topologies of gp120 surface loop that agree with those in reported structural data. Molecular dynamics simulation showed that in the unliganded state, V1/V2 loop settled into a thermodynamically stable arrangement near V3 loop for conformational masking of V3 tip, a potent neutralization epitope. In the CD4-bound state, however, V1/V2 loop was rearranged near the bound CD4 to support CD4 binding. In parallel, cell-based adaptation in the absence of anti-viral antibody pressures led to the identification of amino acid substitutions that individually enhance viral entry and growth efficiencies in association with reduced sensitivity to CCR5 antagonist TAK-779. Notably, all these substitutions were positioned on the receptors binding surfaces in V1/V2 or V3 loop. In silico structural studies predicted some physical changes of gp120 by substitutions with alterations in viral replication phenotypes. These data suggest that V1/V2 loop is critical for creating a gp120 structure that masks co-receptor binding site compatible with maintenance of viral infectivity, and for tuning a functional balance of gp120 between immune escape ability and infectivity to optimize HIV-1 replication fitness.
Keywords: MD simulation; V1/V2 loop; V3 loop; adaptive mutation; homology modeling.
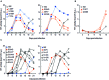
Figures

Similar articles
-
Species-Specific Valid Ternary Interactions of HIV-1 Env-gp120, CD4, and CCR5 as Revealed by an Adaptive Single-Amino Acid Substitution at the V3 Loop Tip.J Virol. 2021 Jun 10;95(13):e0217720. doi: 10.1128/JVI.02177-20. Epub 2021 Jun 10. J Virol. 2021. PMID: 33883222 Free PMC article.
-
Involvement of the V1/V2 variable loop structure in the exposure of human immunodeficiency virus type 1 gp120 epitopes induced by receptor binding.J Virol. 1995 Sep;69(9):5723-33. doi: 10.1128/JVI.69.9.5723-5733.1995. J Virol. 1995. PMID: 7543586 Free PMC article.
-
A novel human antibody against human immunodeficiency virus type 1 gp120 is V1, V2, and V3 loop dependent and helps delimit the epitope of the broadly neutralizing antibody immunoglobulin G1 b12.J Virol. 2003 Jun;77(12):6965-78. doi: 10.1128/jvi.77.12.6965-6978.2003. J Virol. 2003. PMID: 12768015 Free PMC article.
-
Loss of a single N-linked glycan allows CD4-independent human immunodeficiency virus type 1 infection by altering the position of the gp120 V1/V2 variable loops.J Virol. 2001 Apr;75(7):3435-43. doi: 10.1128/JVI.75.7.3435-3443.2001. J Virol. 2001. PMID: 11238869 Free PMC article.
-
V3 loop truncations in HIV-1 envelope impart resistance to coreceptor inhibitors and enhanced sensitivity to neutralizing antibodies.PLoS Pathog. 2007 Aug 24;3(8):e117. doi: 10.1371/journal.ppat.0030117. PLoS Pathog. 2007. PMID: 17722977 Free PMC article.
Cited by
-
Cis-Allosteric Regulation of HIV-1 Reverse Transcriptase by Integrase.Viruses. 2022 Dec 21;15(1):31. doi: 10.3390/v15010031. Viruses. 2022. PMID: 36680070 Free PMC article.
-
Insights into the molecular mechanism underlying CD4-dependency and neutralization sensitivity of HIV-1: a comparative molecular dynamics study on gp120s from isolates with different phenotypes.RSC Adv. 2018 Apr 17;8(26):14355-14368. doi: 10.1039/c8ra00425k. eCollection 2018 Apr 17. RSC Adv. 2018. PMID: 35540760 Free PMC article.
-
Grand Challenge in Human/Animal Virology: Unseen, Smallest Replicative Entities Shape the Whole Globe.Front Microbiol. 2020 Mar 18;11:431. doi: 10.3389/fmicb.2020.00431. eCollection 2020. Front Microbiol. 2020. PMID: 32256480 Free PMC article. No abstract available.
-
Molecular Dynamics Simulation of the Influenza A(H3N2) Hemagglutinin Trimer Reveals the Structural Basis for Adaptive Evolution of the Recent Epidemic Clade 3C.2a.Front Microbiol. 2017 Apr 10;8:584. doi: 10.3389/fmicb.2017.00584. eCollection 2017. Front Microbiol. 2017. PMID: 28443077 Free PMC article.
-
Identification of a Novel Cis-Acting Regulator of HIV-1 Genome Packaging.Int J Mol Sci. 2021 Mar 26;22(7):3435. doi: 10.3390/ijms22073435. Int J Mol Sci. 2021. PMID: 33810482 Free PMC article.
References
-
- Adachi A., Kawamura M., Tokunaga K., Sakai H. (1996). “Methods for HIV/SIV gene analysis,” in Viral Genome Methods ed. Adolph K. W. (Boca Raton, FL: CRC Press; ) 43–53.
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

