Bilateral gene interaction hierarchy analysis of the cell death gene response emphasizes the significance of cell cycle genes following unilateral traumatic brain injury
- PMID: 26912237
- PMCID: PMC4765060
- DOI: 10.1186/s12864-016-2412-0
Bilateral gene interaction hierarchy analysis of the cell death gene response emphasizes the significance of cell cycle genes following unilateral traumatic brain injury
Abstract
Background: Delayed or secondary cell death that is caused by a cascade of cellular and molecular processes initiated by traumatic brain injury (TBI) may be reduced or prevented if an effective neuroprotective strategy is employed. Microarray and subsequent bioinformatic analyses were used to determine which genes, pathways and networks were significantly altered 24 h after unilateral TBI in the rat. Ipsilateral hemi-brain, the corresponding contralateral hemi-brain, and naïve (control) brain tissue were used for microarray analysis.
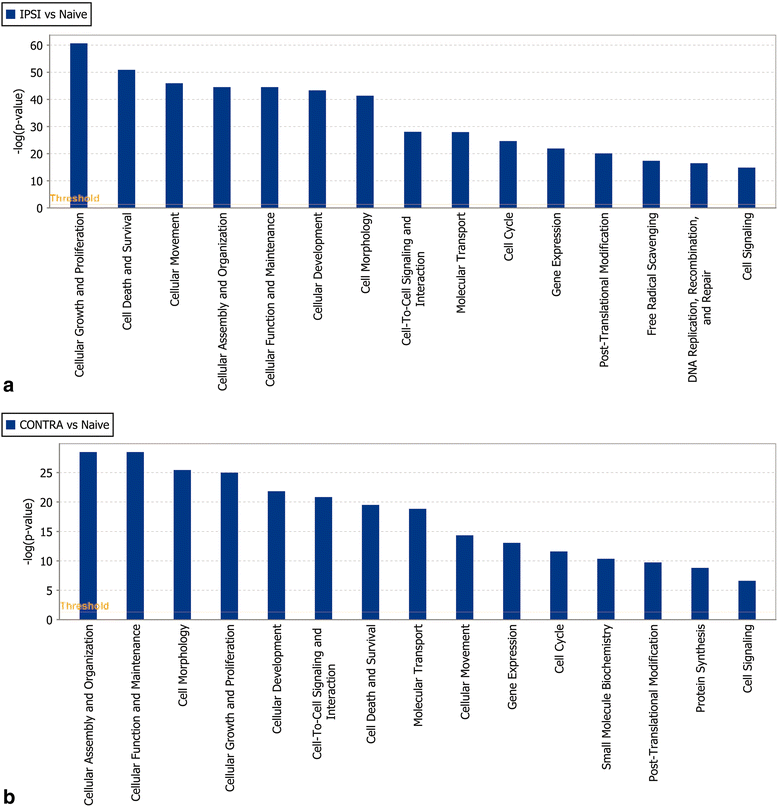
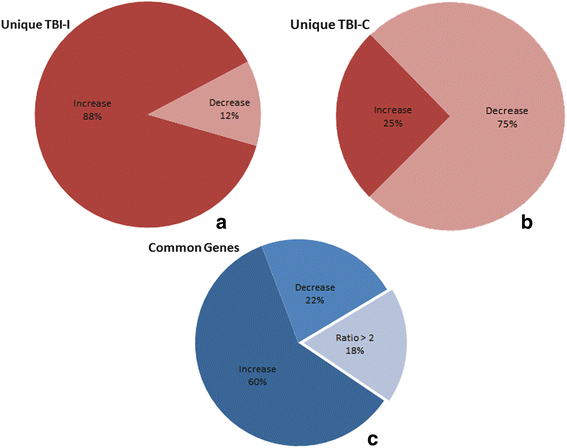
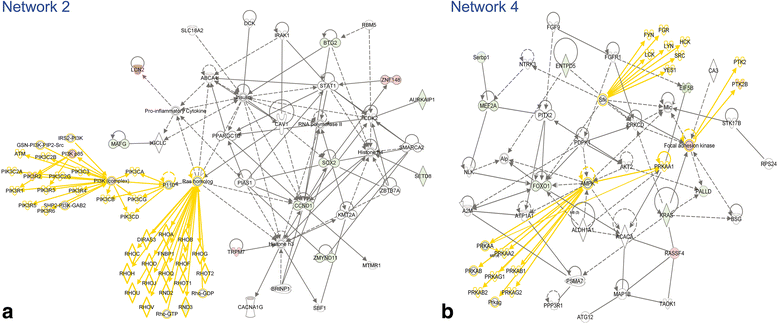
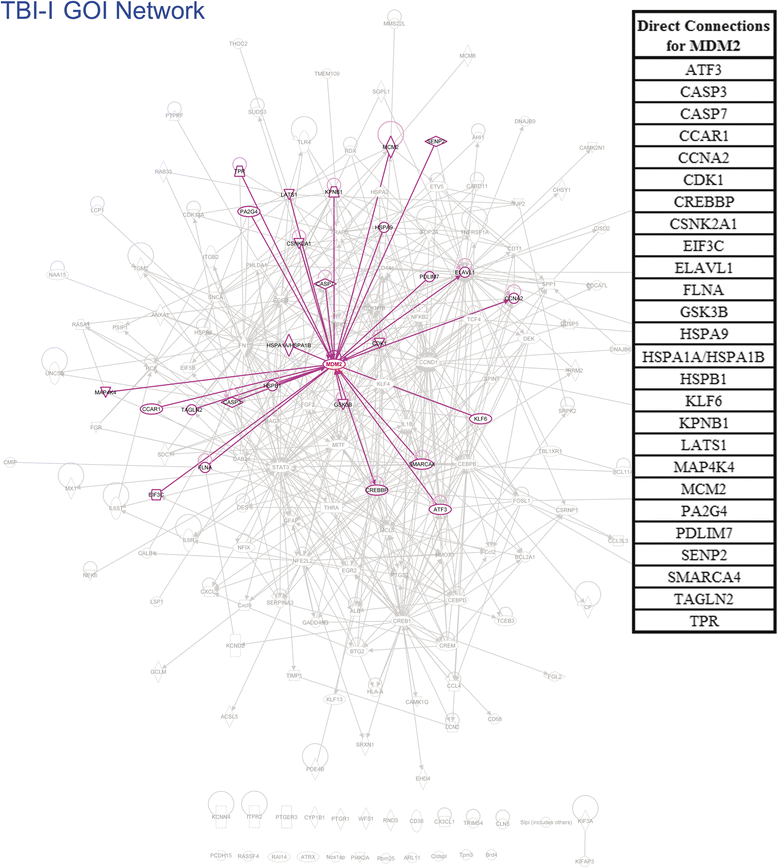
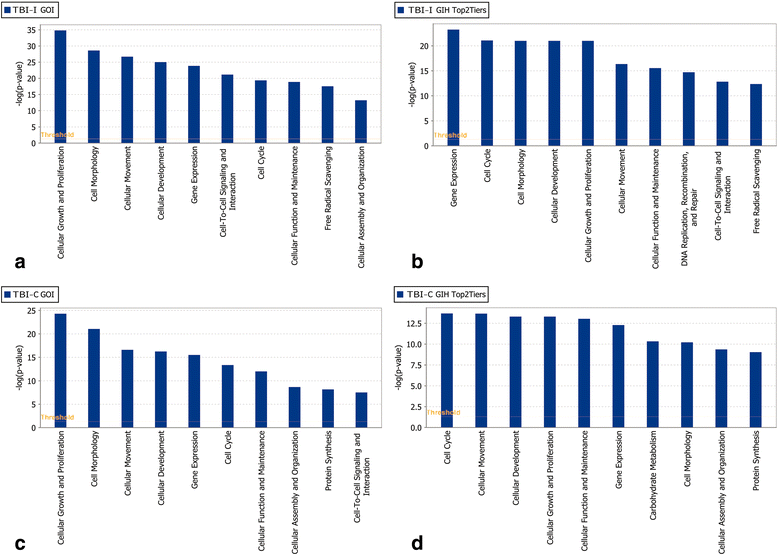
Results: Ingenuity Pathway Analysis showed cell death and survival (CD) to be a top molecular and cellular function associated with TBI on both sides of the brain. One major finding was that the overall gene expression pattern suggested an increase in CD genes in ipsilateral brain tissue and suppression of CD genes contralateral to the injury which may indicate an endogenous protective mechanism. We created networks of genes of interest (GOI) and ranked the genes by the number of direct connections each had in the GOI networks, creating gene interaction hierarchies (GIHs). Cell cycle was determined from the resultant GIHs to be a significant molecular and cellular function in post-TBI CD gene response.
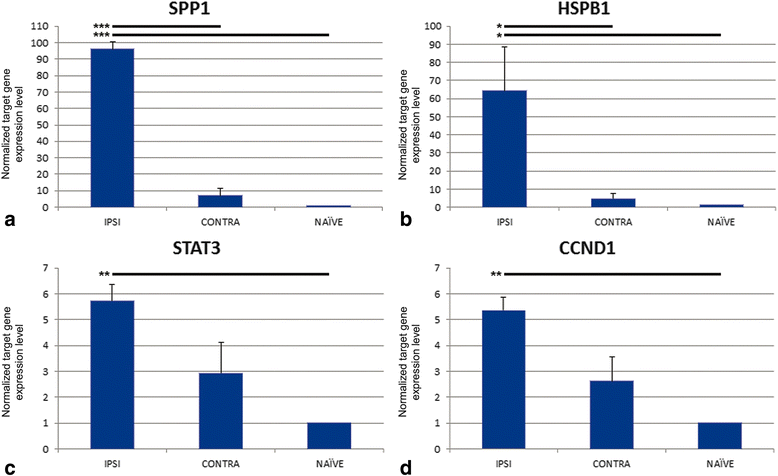
Conclusions: Cell cycle and apoptosis signalling genes that were highly ranked in the GIHs and exhibited either the inverse ipsilateral/contralateral expression pattern or contralateral suppression were identified and included STAT3, CCND1, CCND2, and BAX. Additional exploration into the remote suppression of CD genes may provide insight into neuroprotective mechanisms that could be used to develop therapies to prevent cell death following TBI.
Figures




References
-
- Faul M, Xu L, Wald MM, Coronado VG. Traumatic brain injury in the United States: emergency department visits, hospitalizations and deaths 2002–2006. Atlanta: Centers for Disease Control and Prevention, National Center for Injury Prevention and Control; 2010.
-
- Coronado VG, Xu L, Basavaraju SV, McGuire LC, Wald MM, Faul MD, et al. Surveillance for traumatic brain injury-related deaths--United States, 1997–2007. Morb Mortal Wkly Rep Surveill Summ. 2011;60(5):1–32. - PubMed
-
- Zaloshnja E, Miller T, Langlois JA, Selassie AW. Prevalence of long-term disability from traumatic brain injury in the civilian population of the United States, 2005. J Head Trauma Rehabil. 2008;23(6):394–400. doi: 10.1097/01.HTR.0000341435.52004.ac. - DOI - PubMed
-
- Gubata ME, Packnett ER, Blandford CD, Piccirillo AL, Niebuhr DW, Cowan DN. Trends in the Epidemiology of Disability Related to Traumatic Brain Injury in the US Army and Marine Corps: 2005 to 2010. J Head Trauma Rehabil. 2013 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- T15 LM007056/LM/NLM NIH HHS/United States
- G12 RR003034/RR/NCRR NIH HHS/United States
- U01 NS 057993/NS/NINDS NIH HHS/United States
- KL2 RR025009/RR/NCRR NIH HHS/United States
- S21MD000101/MD/NIMHD NIH HHS/United States
- UL1 TR000454/TR/NCATS NIH HHS/United States
- U54 RR026137/RR/NCRR NIH HHS/United States
- #5P20M006131-02/PHS HHS/United States
- UL1 RR025008/RR/NCRR NIH HHS/United States
- G12RR003034/RR/NCRR NIH HHS/United States
- U54 MD007588/MD/NIMHD NIH HHS/United States
- KL2 TR000455/TR/NCATS NIH HHS/United States
- #52006306/HHMI/Howard Hughes Medical Institute/United States
- TL1 RR025010/RR/NCRR NIH HHS/United States
- U01 NS057993/NS/NINDS NIH HHS/United States
- RR025009/RR/NCRR NIH HHS/United States
- S21 MD000101/MD/NIMHD NIH HHS/United States
- U54 NS060659/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

