A polyalanine peptide derived from polar fish with anti-infectious activities
- PMID: 26916401
- PMCID: PMC4768251
- DOI: 10.1038/srep21385
A polyalanine peptide derived from polar fish with anti-infectious activities
Erratum in
-
Corrigendum: A polyalanine peptide derived from polar fish with anti-infectious activities.Sci Rep. 2016 Jul 20;6:28216. doi: 10.1038/srep28216. Sci Rep. 2016. PMID: 27435893 Free PMC article. No abstract available.
Abstract
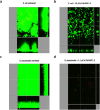
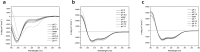
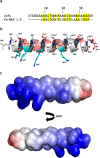
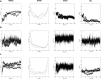
Due to the growing concern about antibiotic-resistant microbial infections, increasing support has been given to new drug discovery programs. A promising alternative to counter bacterial infections includes the antimicrobial peptides (AMPs), which have emerged as model molecules for rational design strategies. Here we focused on the study of Pa-MAP 1.9, a rationally designed AMP derived from the polar fish Pleuronectes americanus. Pa-MAP 1.9 was active against Gram-negative planktonic bacteria and biofilms, without being cytotoxic to mammalian cells. By using AFM, leakage assays, CD spectroscopy and in silico tools, we found that Pa-MAP 1.9 may be acting both on intracellular targets and on the bacterial surface, also being more efficient at interacting with anionic LUVs mimicking Gram-negative bacterial surface, where this peptide adopts α-helical conformations, than cholesterol-enriched LUVs mimicking mammalian cells. Thus, as bacteria present varied physiological features that favor antibiotic-resistance, Pa-MAP 1.9 could be a promising candidate in the development of tools against infections caused by pathogenic bacteria.
Figures


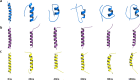

Similar articles
-
Anti-Protozoan Activities of Polar Fish-Derived Polyalanine Synthetic Peptides.Mar Drugs. 2023 Jul 31;21(8):434. doi: 10.3390/md21080434. Mar Drugs. 2023. PMID: 37623715 Free PMC article.
-
In vivo antimicrobial evaluation of an alanine-rich peptide derived from Pleuronectes americanus.Peptides. 2013 Apr;42:144-8. doi: 10.1016/j.peptides.2013.02.001. Epub 2013 Feb 13. Peptides. 2013. PMID: 23416023
-
Structural and functional evaluation of the palindromic alanine-rich antimicrobial peptide Pa-MAP2.Biochim Biophys Acta. 2016 Jul;1858(7 Pt A):1488-98. doi: 10.1016/j.bbamem.2016.04.003. Epub 2016 Apr 8. Biochim Biophys Acta. 2016. PMID: 27063608
-
Antimicrobial peptides as potential anti-biofilm agents against multidrug-resistant bacteria.J Microbiol Immunol Infect. 2017 Aug;50(4):405-410. doi: 10.1016/j.jmii.2016.12.005. Epub 2017 Jun 26. J Microbiol Immunol Infect. 2017. PMID: 28690026 Review.
-
Antimicrobial peptides: new candidates in the fight against bacterial infections.Biopolymers. 2005;80(6):717-35. doi: 10.1002/bip.20286. Biopolymers. 2005. PMID: 15880793 Review.
Cited by
-
Structure-function-guided exploration of the antimicrobial peptide polybia-CP identifies activity determinants and generates synthetic therapeutic candidates.Commun Biol. 2018 Dec 7;1:221. doi: 10.1038/s42003-018-0224-2. eCollection 2018. Commun Biol. 2018. PMID: 30534613 Free PMC article.
-
Anti-Protozoan Activities of Polar Fish-Derived Polyalanine Synthetic Peptides.Mar Drugs. 2023 Jul 31;21(8):434. doi: 10.3390/md21080434. Mar Drugs. 2023. PMID: 37623715 Free PMC article.
-
Antibiofilm Activity of Acidic Phospholipase Isoform Isolated from Bothrops erythromelas Snake Venom.Toxins (Basel). 2020 Sep 20;12(9):606. doi: 10.3390/toxins12090606. Toxins (Basel). 2020. PMID: 32962193 Free PMC article.
-
Combating Antimicrobial Resistance With New-To-Nature Lanthipeptides Created by Genetic Code Expansion.Front Microbiol. 2020 Nov 5;11:590522. doi: 10.3389/fmicb.2020.590522. eCollection 2020. Front Microbiol. 2020. PMID: 33250877 Free PMC article. Review.
-
Adevonin, a novel synthetic antimicrobial peptide designed from the Adenanthera pavonina trypsin inhibitor (ApTI) sequence.Pathog Glob Health. 2018 Dec;112(8):438-447. doi: 10.1080/20477724.2018.1559489. Epub 2018 Dec 20. Pathog Glob Health. 2018. PMID: 30570384 Free PMC article.
References
-
- Alanis A. J. Resistance to antibiotics: Are we in the post-antibiotic era? Arch. Med. Res. 36, 697–705 (2005). - PubMed
-
- Giamarellou H. & Poulakou G. Multidrug-Resistant Gram-Negative Infections What are the Treatment Options? Drugs 69, 1879–1901 (2009). - PubMed
-
- de la Fuente-Núñez C., Reffuveille F., Fernandez L. & Hancock R. E. W. Bacterial biofilm development as a multicellular adaptation: antibiotic resistance and new therapeutic strategies. Curr. Opin. Microbiol. 16, 580–589 (2013). - PubMed
-
- Van Acker H., Van Dijck P. & Coenye T. Molecular mechanisms of antimicrobial tolerance and resistance in bacterial and fungal biofilms. Trends Microbiol. 22, 326–333 (2014). - PubMed
-
- Fjell C. D., Hiss J. A., Hancock R. E. & Schneider G. Designing antimicrobial peptides: form follows function. Nat. Rev. Drug. Discov. 11, 37–51 (2012). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

