Synthesis and evaluation of an alkyne-modified ATP analog for enzymatic incorporation into RNA
- PMID: 26927424
- PMCID: PMC4785081
- DOI: 10.1016/j.bmcl.2016.02.038
Synthesis and evaluation of an alkyne-modified ATP analog for enzymatic incorporation into RNA
Abstract
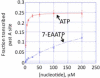
Alkyne-modified nucleoside analogs are useful for nucleic acid localization as well as functional and structural studies because of their ability to participate in copper-catalyzed azide/alkyne cycloaddition (CuAAC) reactions. Here we describe the synthesis of the triphosphate of 7-ethynyl-8-aza-7-deazaadenosine (7-EAATP) and the enzymatic incorporation of 7-EAA into RNA. The free nucleoside of 7-EAA is taken up by HeLa cells and incorporated into cellular RNA by endogenous RNA polymerases. In addition, 7-EAATP is a substrate for both T7 RNA polymerase and poly (A) polymerase from Escherichia coli in vitro, albeit at lower efficiencies than with ATP. This work adds to the toolbox of nucleoside analogs useful for RNA labeling.
Keywords: ATP analogs; Cellular RNA labeling; Click chemistry; Polyadenylation; T7 transcription.
Copyright © 2016 Elsevier Ltd. All rights reserved.
Figures



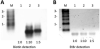
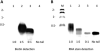
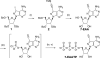
Similar articles
-
Synthesis of 2-Substituted Adenosine Triphosphate Derivatives and their use in Enzymatic Synthesis and Postsynthetic Labelling of RNA.Chembiochem. 2025 Jun 16;26(12):e202500241. doi: 10.1002/cbic.202500241. Epub 2025 May 21. Chembiochem. 2025. PMID: 40317833 Free PMC article.
-
Click chemistry for rapid labeling and ligation of RNA.Chembiochem. 2011 Jan 3;12(1):125-31. doi: 10.1002/cbic.201000466. Chembiochem. 2011. PMID: 21132831
-
Chemical reporters for monitoring RNA synthesis and poly(A) tail dynamics.Chembiochem. 2012 May 29;13(8):1112-5. doi: 10.1002/cbic.201200091. Epub 2012 Apr 19. Chembiochem. 2012. PMID: 22513998 Free PMC article.
-
Advances in Evolved T7 RNA Polymerases for Expanding the Frontiers of Enzymatic Nucleic Acid Synthesis.Chembiochem. 2024 Dec 16;25(24):e202400483. doi: 10.1002/cbic.202400483. Epub 2024 Sep 23. Chembiochem. 2024. PMID: 39085046 Review.
-
Azide-Modified Nucleosides as Versatile Tools for Bioorthogonal Labeling and Functionalization.Chem Rec. 2022 May;22(5):e202100322. doi: 10.1002/tcr.202100322. Epub 2022 Feb 21. Chem Rec. 2022. PMID: 35189013 Review.
Cited by
-
Exploiting Endogenous Enzymes for Cancer-Cell Selective Metabolic Labeling of RNA in Vivo.J Am Chem Soc. 2022 Apr 27;144(16):7085-7088. doi: 10.1021/jacs.2c02404. Epub 2022 Apr 13. J Am Chem Soc. 2022. PMID: 35416650 Free PMC article.
-
Nucleoside analogs in the study of the epitranscriptome.Methods. 2019 Mar 1;156:46-52. doi: 10.1016/j.ymeth.2018.10.014. Epub 2018 Oct 26. Methods. 2019. PMID: 30827466 Free PMC article. Review.
-
Post-transcriptional labeling by using Suzuki-Miyaura cross-coupling generates functional RNA probes.Nucleic Acids Res. 2018 Jun 20;46(11):e65. doi: 10.1093/nar/gky185. Nucleic Acids Res. 2018. PMID: 29546376 Free PMC article.
-
Modified nucleoside triphosphates in bacterial research for in vitro and live-cell applications.RSC Chem Biol. 2020 Dec 1;1(5):333-351. doi: 10.1039/d0cb00078g. Epub 2020 Sep 14. RSC Chem Biol. 2020. PMID: 33928252 Free PMC article.
-
An optimized chemical-genetic method for cell-specific metabolic labeling of RNA.Nat Methods. 2020 Mar;17(3):311-318. doi: 10.1038/s41592-019-0726-y. Epub 2020 Feb 3. Nat Methods. 2020. PMID: 32015544 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

