A Hybrid of the Chemical Master Equation and the Gillespie Algorithm for Efficient Stochastic Simulations of Sub-Networks
- PMID: 26930199
- PMCID: PMC4773016
- DOI: 10.1371/journal.pone.0149909
A Hybrid of the Chemical Master Equation and the Gillespie Algorithm for Efficient Stochastic Simulations of Sub-Networks
Abstract
Modeling stochastic behavior of chemical reaction networks is an important endeavor in many aspects of chemistry and systems biology. The chemical master equation (CME) and the Gillespie algorithm (GA) are the two most fundamental approaches to such modeling; however, each of them has its own limitations: the GA may require long computing times, while the CME may demand unrealistic memory storage capacity. We propose a method that combines the CME and the GA that allows one to simulate stochastically a part of a reaction network. First, a reaction network is divided into two parts. The first part is simulated via the GA, while the solution of the CME for the second part is fed into the GA in order to update its propensities. The advantage of this method is that it avoids the need to solve the CME or stochastically simulate the entire network, which makes it highly efficient. One of its drawbacks, however, is that most of the information about the second part of the network is lost in the process. Therefore, this method is most useful when only partial information about a reaction network is needed. We tested this method against the GA on two systems of interest in biology--the gene switch and the Griffith model of a genetic oscillator--and have shown it to be highly accurate. Comparing this method to four different stochastic algorithms revealed it to be at least an order of magnitude faster than the fastest among them.
Conflict of interest statement
Figures
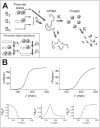



Similar articles
-
Stochastic modeling and numerical simulation of gene regulatory networks with protein bursting.J Theor Biol. 2017 May 21;421:51-70. doi: 10.1016/j.jtbi.2017.03.017. Epub 2017 Mar 21. J Theor Biol. 2017. PMID: 28341132
-
Distributions for negative-feedback-regulated stochastic gene expression: dimension reduction and numerical solution of the chemical master equation.J Theor Biol. 2010 May 21;264(2):377-85. doi: 10.1016/j.jtbi.2010.02.004. Epub 2010 Feb 6. J Theor Biol. 2010. PMID: 20144620
-
Periodic synchronization of isolated network elements facilitates simulating and inferring gene regulatory networks including stochastic molecular kinetics.BMC Bioinformatics. 2022 Jan 5;23(1):13. doi: 10.1186/s12859-021-04541-6. BMC Bioinformatics. 2022. PMID: 34986805 Free PMC article.
-
Stochastic approaches in systems biology.Wiley Interdiscip Rev Syst Biol Med. 2010 Jul-Aug;2(4):385-397. doi: 10.1002/wsbm.78. Wiley Interdiscip Rev Syst Biol Med. 2010. PMID: 20836037 Review.
-
Exact and approximate stochastic simulation of intracellular calcium dynamics.J Biomed Biotechnol. 2011;2011:572492. doi: 10.1155/2011/572492. Epub 2011 Nov 9. J Biomed Biotechnol. 2011. PMID: 22131814 Free PMC article. Review.
Cited by
-
Modelling gambiense human African trypanosomiasis infection in villages of the Democratic Republic of Congo using Kolmogorov forward equations.J R Soc Interface. 2021 Oct;18(183):20210419. doi: 10.1098/rsif.2021.0419. Epub 2021 Oct 6. J R Soc Interface. 2021. PMID: 34610258 Free PMC article.
-
Path integral approach to generating functions for multistep post-transcription and post-translation processes and arbitrary initial conditions.J Math Biol. 2019 Dec;79(6-7):2211-2236. doi: 10.1007/s00285-019-01426-4. Epub 2019 Sep 5. J Math Biol. 2019. PMID: 31511967
-
Exploration of Serum Proteomic Profiling and Diagnostic Model That Differentiate Crohn's Disease and Intestinal Tuberculosis.PLoS One. 2016 Dec 20;11(12):e0167109. doi: 10.1371/journal.pone.0167109. eCollection 2016. PLoS One. 2016. PMID: 27997555 Free PMC article. Clinical Trial.
References
-
- Doob JL. Topics in the Theory of Markoff Chains. Transactions of the American Mathematical Society 1945; 58 (3), 455–473 10.2307/1990339 - DOI
-
- Gillespie DT. Exact Stochastic Simulation of Coupled Chemical Reactions. J. Phys. Chem. 1977; 81(25), 2340–2361 10.1021/j100540a008 - DOI
-
- Gibson MA, Bruck J. Efficient Exact Stochastic Simulation of Chemical Systems with Many Species and Many Channels. J. Phys. Chem. 2000; 104(9), 1876–1889 10.1021/jp993732q - DOI
-
- Cao Y, Li H, Petzold L. Efficient formulation of the stochastic simulation algorithm for chemically reacting systems. J. Chem. Phys. 2004; 121, 4059 - PubMed
-
- Gillespie DT. Approximate accelerated stochastic simulation of chemically reacting systems. J. Chem. Phys. 2001; 115(4), 1716 10.1063/1.1378322 - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

