Interaction of Mycobacterium tuberculosis Virulence Factor RipA with Chaperone MoxR1 Is Required for Transport through the TAT Secretion System
- PMID: 26933057
- PMCID: PMC4810496
- DOI: 10.1128/mBio.02259-15
Interaction of Mycobacterium tuberculosis Virulence Factor RipA with Chaperone MoxR1 Is Required for Transport through the TAT Secretion System
Abstract
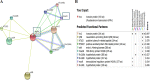
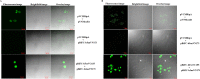
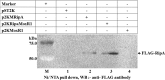
Mycobacterium tuberculosis is a leading cause of death worldwide. The M. tuberculosis TAT (twin-arginine translocation) protein secretion system is present at the cytoplasmic membrane of mycobacteria and is known to transport folded proteins. The TAT secretion system is reported to be essential for many important bacterial processes that include cell wall biosynthesis. The M. tuberculosis secretion and invasion protein RipA has endopeptidase activity and interacts with one of the resuscitation antigens (RpfB) that are expressed during pathogen reactivation. MoxR1, a member of the ATPase family that is associated with various cellular activities, was predicted to interact with RipA based on in silico analyses. A bimolecular fluorescence complementation (BiFC) assay confirmed the interaction of these two proteins in HEK293T cells. The overexpression of RipA in Mycobacterium smegmatis and copurification with MoxR1 further validated their interaction in vivo. Recombinant MoxR1 protein, expressed in Escherichia coli, displays ATP-enhanced chaperone activity. Secretion of recombinant RipA (rRipA) protein into the E. coli culture filtrate was not observed in the absence of RipA-MoxR interaction. Inhibition of this export system in M. tuberculosis, including the key players, will prevent localization of peptidoglycan hydrolase and result in sensitivity to existing β-lactam antibiotics, opening up new candidates for drug repurposing.
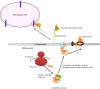
Importance: The virulence mechanism of mycobacteria is very complex. Broadly, the virulence factors can be classified as secretion factors, cell surface components, enzymes involved in cellular metabolism, and transcriptional regulators. The mycobacteria have evolved several mechanisms to secrete its proteins. Here, we have identified one of the virulence proteins of Mycobacterium tuberculosis, RipA, possessing peptidoglycan hydrolase activities secreted by the TAT secretion pathway. We also identified MoxR1 as a protein-protein interaction partner of RipA and demonstrated chaperone activity of this protein. We show that MoxR1-mediated folding is critical for the secretion of RipA within the TAT system. Inhibition of this export system in M. tuberculosis will prevent localization of peptidoglycan hydrolase and result in sensitivity to existing β-lactam antibiotics, opening up new candidates for drug repurposing.
Copyright © 2016 Bhuwan et al.
Figures

Similar articles
-
The twin-arginine translocation pathway of Mycobacterium smegmatis is functional and required for the export of mycobacterial beta-lactamases.J Bacteriol. 2005 Nov;187(22):7667-79. doi: 10.1128/JB.187.22.7667-7679.2005. J Bacteriol. 2005. PMID: 16267291 Free PMC article.
-
A partner for the resuscitation-promoting factors of Mycobacterium tuberculosis.Mol Microbiol. 2007 Nov;66(3):658-68. doi: 10.1111/j.1365-2958.2007.05945.x. Epub 2007 Oct 4. Mol Microbiol. 2007. PMID: 17919286
-
A protein secretion pathway critical for Mycobacterium tuberculosis virulence is conserved and functional in Mycobacterium smegmatis.J Bacteriol. 2005 Feb;187(4):1238-45. doi: 10.1128/JB.187.4.1238-1245.2005. J Bacteriol. 2005. PMID: 15687187 Free PMC article.
-
Protein export systems of Mycobacterium tuberculosis: novel targets for drug development?Future Microbiol. 2010 Oct;5(10):1581-97. doi: 10.2217/fmb.10.112. Future Microbiol. 2010. PMID: 21073315 Free PMC article. Review.
-
Lipid transport in Mycobacterium tuberculosis and its implications in virulence and drug development.Biochem Pharmacol. 2015 Aug 1;96(3):159-67. doi: 10.1016/j.bcp.2015.05.001. Epub 2015 May 16. Biochem Pharmacol. 2015. PMID: 25986884 Review.
Cited by
-
Mycobacterium tuberculosis RipA Dampens TLR4-Mediated Host Protective Response Using a Multi-Pronged Approach Involving Autophagy, Apoptosis, Metabolic Repurposing, and Immune Modulation.Front Immunol. 2021 Mar 4;12:636644. doi: 10.3389/fimmu.2021.636644. eCollection 2021. Front Immunol. 2021. PMID: 33746976 Free PMC article.
-
Tat-exported peptidoglycan amidase-dependent cell division contributes to Salmonella Typhimurium fitness in the inflamed gut.PLoS Pathog. 2018 Oct 31;14(10):e1007391. doi: 10.1371/journal.ppat.1007391. eCollection 2018 Oct. PLoS Pathog. 2018. PMID: 30379938 Free PMC article.
-
Not just an antibiotic target: Exploring the role of type I signal peptidase in bacterial virulence.Bioorg Med Chem. 2016 Dec 15;24(24):6370-6378. doi: 10.1016/j.bmc.2016.09.048. Epub 2016 Sep 21. Bioorg Med Chem. 2016. PMID: 27769673 Free PMC article.
-
The Next Frontier in Tuberculosis Investigation: Automated Whole Genome Sequencing for Mycobacterium tuberculosis Analysis.Int J Mol Sci. 2024 Jul 19;25(14):7909. doi: 10.3390/ijms25147909. Int J Mol Sci. 2024. PMID: 39063151 Free PMC article.
-
Silver Nanoparticles for the Therapy of Tuberculosis.Int J Nanomedicine. 2020 Mar 31;15:2231-2258. doi: 10.2147/IJN.S241183. eCollection 2020. Int J Nanomedicine. 2020. PMID: 32280217 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

