The impact of structural genomics: the first quindecennial
- PMID: 26935210
- PMCID: PMC4834271
- DOI: 10.1007/s10969-016-9201-5
The impact of structural genomics: the first quindecennial
Abstract
The period 2000-2015 brought the advent of high-throughput approaches to protein structure determination. With the overall funding on the order of $2 billion (in 2010 dollars), the structural genomics (SG) consortia established worldwide have developed pipelines for target selection, protein production, sample preparation, crystallization, and structure determination by X-ray crystallography and NMR. These efforts resulted in the determination of over 13,500 protein structures, mostly from unique protein families, and increased the structural coverage of the expanding protein universe. SG programs contributed over 4400 publications to the scientific literature. The NIH-funded Protein Structure Initiatives alone have produced over 2000 scientific publications, which to date have attracted more than 93,000 citations. Software and database developments that were necessary to handle high-throughput structure determination workflows have led to structures of better quality and improved integrity of the associated data. Organized and accessible data have a positive impact on the reproducibility of scientific experiments. Most of the experimental data generated by the SG centers are freely available to the community and has been utilized by scientists in various fields of research. SG projects have created, improved, streamlined, and validated many protocols for protein production and crystallization, data collection, and functional analysis, significantly benefiting biological and biomedical research.
Keywords: NMR; Protein crystallography; Structural genomics.
Figures
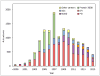
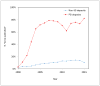




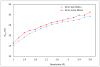
References
-
- NIGMS. [Accessed November 4, 2015];PSI Pilot Phase Fact Sheet. 2001 https://www.nigms.nih.gov/Research/specificareas/PSI/background/Pages/Pi....
Publication types
MeSH terms
Substances
Grants and funding
- GM093324/GM/NIGMS NIH HHS/United States
- HHSN272201200026C/PHS HHS/United States
- GM094662/GM/NIGMS NIH HHS/United States
- HG008424/HG/NHGRI NIH HHS/United States
- U01 HG008424/HG/NHGRI NIH HHS/United States
- U54 GM094585/GM/NIGMS NIH HHS/United States
- GM094585/GM/NIGMS NIH HHS/United States
- GM093342/GM/NIGMS NIH HHS/United States
- U54 GM094662/GM/NIGMS NIH HHS/United States
- U54 GM093342/GM/NIGMS NIH HHS/United States
- U01 GM093324/GM/NIGMS NIH HHS/United States
- HHSN272201200026C/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources

