CHIR99021 enhances Klf4 Expression through β-Catenin Signaling and miR-7a Regulation in J1 Mouse Embryonic Stem Cells
- PMID: 26938105
- PMCID: PMC4777400
- DOI: 10.1371/journal.pone.0150936
CHIR99021 enhances Klf4 Expression through β-Catenin Signaling and miR-7a Regulation in J1 Mouse Embryonic Stem Cells
Abstract
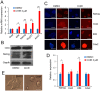
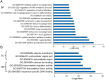
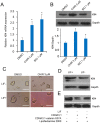
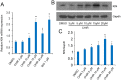
Understanding the mechanisms that regulate pluripotency of embryonic stem cells (ESCs) is important to ensure their safe clinical use. CHIR99021 (CHIR)-induced activation of Wnt/β-catenin signaling promotes self-renewal in mouse ESCs (mESCs). β-catenin functions individually or cooperates with transcription factors to activate stemness factors such as c-Myc, Esrrb, Pou5f1, and Nanog. However the relationship between the core pluripotent factor, Kruppel-like factor 4 (also known as GKLF or EZF) and Wnt/β-catenin signaling, remains ambiguous in J1 mESCs. DNA microarray analysis revealed that CHIR-treatment promoted pluripotency-maintaining transcription factors and repressed germ layer specification markers. CHIR also promoted genes related to the development of extracellular regions and the plasma membrane to maintain pluripotency of J1 mESCs. Among the CHIR-regulated genes, Klf4 has not been reported previously. We identified a novel cis element in the Klf4 gene that was activated by β-catenin in J1 mESCs. We determined that β-catenin interacted with this cis element, identifying Klf4 as a β-catenin target gene in this context. Moreover, several microRNAs that targeted the 3'-UTR of Klf4 mRNA were identified, with miR-7a being down-regulated by CHIR in a β-catenin-independent manner in J1 mESCs. These data collectively suggest that CHIR enhances Klf4 expression by repressing miR-7a expression or canonical Wnt pathway activation.
Conflict of interest statement
Figures

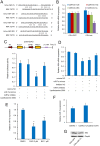
Similar articles
-
GSK3 inhibitors CHIR99021 and 6-bromoindirubin-3'-oxime inhibit microRNA maturation in mouse embryonic stem cells.Sci Rep. 2015 Mar 2;5:8666. doi: 10.1038/srep08666. Sci Rep. 2015. PMID: 25727520 Free PMC article.
-
A feedback loop comprising PRMT7 and miR-24-2 interplays with Oct4, Nanog, Klf4 and c-Myc to regulate stemness.Nucleic Acids Res. 2016 Dec 15;44(22):10603-10618. doi: 10.1093/nar/gkw788. Epub 2016 Sep 12. Nucleic Acids Res. 2016. PMID: 27625395 Free PMC article.
-
CHIR99021 promotes self-renewal of mouse embryonic stem cells by modulation of protein-encoding gene and long intergenic non-coding RNA expression.Exp Cell Res. 2013 Oct 15;319(17):2684-99. doi: 10.1016/j.yexcr.2013.08.027. Epub 2013 Sep 7. Exp Cell Res. 2013. PMID: 24021571
-
The genetics of induced pluripotency.Reproduction. 2010 Jan;139(1):35-44. doi: 10.1530/REP-09-0024. Reproduction. 2010. PMID: 19605512 Review.
-
Wnt/beta-catenin signaling: a promising new target for fibrosis diseases.Physiol Res. 2012;61(4):337-46. doi: 10.33549/physiolres.932289. Epub 2012 Jun 6. Physiol Res. 2012. PMID: 22670697 Review.
Cited by
-
The concurrent stimulation of Wnt and FGF8 signaling induce differentiation of dental mesenchymal cells into odontoblast-like cells.Med Mol Morphol. 2022 Mar;55(1):8-19. doi: 10.1007/s00795-021-00297-3. Epub 2021 Nov 5. Med Mol Morphol. 2022. PMID: 34739612 Free PMC article.
-
Characterization of Stem-like Circulating Tumor Cells in Pancreatic Cancer.Diagnostics (Basel). 2020 May 15;10(5):305. doi: 10.3390/diagnostics10050305. Diagnostics (Basel). 2020. PMID: 32429174 Free PMC article.
-
Negative Regulation of Kruppel-Like Factor 4 on microRNA-106a at Upstream Transcriptional Level and the Role in Gastric Cancer Metastasis.Dig Dis Sci. 2018 Oct;63(10):2604-2616. doi: 10.1007/s10620-018-5143-z. Epub 2018 Jun 8. Dig Dis Sci. 2018. PMID: 29948558
-
Modulation of Wnt Signaling Enhances Inner Ear Organoid Development in 3D Culture.PLoS One. 2016 Sep 8;11(9):e0162508. doi: 10.1371/journal.pone.0162508. eCollection 2016. PLoS One. 2016. PMID: 27607106 Free PMC article.
-
Retinoic Acid Induces Differentiation of Mouse F9 Embryonic Carcinoma Cell by Modulating the miR-485 Targeting of Abhd2.Int J Mol Sci. 2019 Apr 26;20(9):2071. doi: 10.3390/ijms20092071. Int J Mol Sci. 2019. PMID: 31035455 Free PMC article.
References
-
- Evans MJ, Kaufman MH. Establishment in culture of pluripotential cells from mouse embryos. Nature. 1981;292(5819):154–6. . - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

