Physiological Properties and Genome Structure of the Hyperthermophilic Filamentous Phage φOH3 Which Infects Thermus thermophilus HB8
- PMID: 26941711
- PMCID: PMC4763002
- DOI: 10.3389/fmicb.2016.00050
Physiological Properties and Genome Structure of the Hyperthermophilic Filamentous Phage φOH3 Which Infects Thermus thermophilus HB8
Abstract
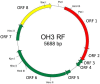
A filamentous bacteriophage, φOH3, was isolated from hot spring sediment in Obama hot spring in Japan with the hyperthermophilic bacterium Thermus thermophilus HB8 as its host. Phage φOH3, which was classified into the Inoviridae family, consists of a flexible filamentous particle 830 nm long and 8 nm wide. φOH3 was stable at temperatures ranging from 70 to 90°C and at pHs ranging from 6 to 9. A one-step growth curve of the phage showed a 60-min latent period beginning immediately postinfection, followed by intracellular virus particle production during the subsequent 40 min. The released virion number of φOH3 was 109. During the latent period, both single stranded DNA (ssDNA) and the replicative form (RF) of phage DNA were multiplied from min 40 onward. During the release period, the copy numbers of both ssDNA and RF DNA increased sharply. The size of the φOH3 genome is 5688 bp, and eight putative open reading frames (ORFs) were annotated. These ORFs were encoded on the plus strand of RF DNA and showed no significant homology with any known phage genes, except ORF 5, which showed 60% identity with the gene VIII product of the Thermus filamentous phage PH75. All the ORFs were similar to predicted genes annotated in the Thermus aquaticus Y51MC23 and Meiothermus timidus DSM 17022 genomes at the amino acid sequence level. This is the first report of the whole genome structure and DNA multiplication of a filamentous T. thermophilus phage within its host cell.
Keywords: Inoviridae; Thermus thermophilus; filamentous phage; hyperthermophilic phage; replicative form.
Figures







Similar articles
-
Structural characterization of the filamentous bacteriophage PH75 from Thermus thermophilus by Raman and UV-resonance Raman spectroscopy.Biochemistry. 2005 Mar 1;44(8):3091-100. doi: 10.1021/bi048163d. Biochemistry. 2005. PMID: 15723554
-
Filamentous phage active on the gram-positive bacterium Propionibacterium freudenreichii.J Bacteriol. 2002 Apr;184(7):2030-3. doi: 10.1128/JB.184.7.2030-2033.2002. J Bacteriol. 2002. PMID: 11889111 Free PMC article.
-
Isolation and genomic analysis of a type IV pili-independent Thermus thermophilus phage, φMN1 from a Japanese hot spring.J Gen Appl Microbiol. 2023 Nov 15;69(2):117-124. doi: 10.2323/jgam.2023.06.008. Epub 2023 Jul 7. J Gen Appl Microbiol. 2023. PMID: 37423744
-
Thermostable repair enzyme for oxidative DNA damage from extremely thermophilic bacterium, Thermus thermophilus HB8.Nucleic Acids Res. 1998 Feb 15;26(4):903-10. doi: 10.1093/nar/26.4.903. Nucleic Acids Res. 1998. PMID: 9461446 Free PMC article.
-
Chemical strategies for the covalent modification of filamentous phage.Front Microbiol. 2014 Dec 23;5:734. doi: 10.3389/fmicb.2014.00734. eCollection 2014. Front Microbiol. 2014. PMID: 25566240 Free PMC article. Review.
Cited by
-
Biogeography and taxonomic overview of terrestrial hot spring thermophilic phages.Extremophiles. 2018 Nov;22(6):827-837. doi: 10.1007/s00792-018-1052-5. Epub 2018 Aug 18. Extremophiles. 2018. PMID: 30121708 Review.
-
Natural diversity of CRISPR spacers of Thermus: evidence of local spacer acquisition and global spacer exchange.Philos Trans R Soc Lond B Biol Sci. 2019 May 13;374(1772):20180092. doi: 10.1098/rstb.2018.0092. Philos Trans R Soc Lond B Biol Sci. 2019. PMID: 30905291 Free PMC article.
-
Bacteriophages of Thermophilic 'Bacillus Group' Bacteria-A Review.Microorganisms. 2021 Jul 16;9(7):1522. doi: 10.3390/microorganisms9071522. Microorganisms. 2021. PMID: 34361957 Free PMC article. Review.
-
The Diversity of Bacteriophages in Hot Springs.Methods Mol Biol. 2024;2738:73-88. doi: 10.1007/978-1-0716-3549-0_4. Methods Mol Biol. 2024. PMID: 37966592 Review.
-
Isolation and characterization of two homolog phages infecting Pseudomonas aeruginosa.Front Microbiol. 2022 Jul 14;13:946251. doi: 10.3389/fmicb.2022.946251. eCollection 2022. Front Microbiol. 2022. PMID: 35935197 Free PMC article.
References
-
- Adams M. H., Anderson E., Kellenberger E. (1959). Bacteriophages. New York, NY: Interscience publishers.
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

