CYP96T1 of Narcissus sp. aff. pseudonarcissus Catalyzes Formation of the Para-Para' C-C Phenol Couple in the Amaryllidaceae Alkaloids
- PMID: 26941773
- PMCID: PMC4766306
- DOI: 10.3389/fpls.2016.00225
CYP96T1 of Narcissus sp. aff. pseudonarcissus Catalyzes Formation of the Para-Para' C-C Phenol Couple in the Amaryllidaceae Alkaloids
Abstract
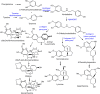
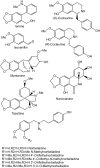
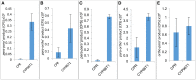
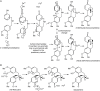
The Amaryllidaceae alkaloids are a family of amino acid derived alkaloids with many biological activities; examples include haemanthamine, haemanthidine, galanthamine, lycorine, and maritidine. Central to the biosynthesis of the majority of these alkaloids is a C-C phenol-coupling reaction that can have para-para', para-ortho', or ortho-para' regiospecificity. Through comparative transcriptomics of Narcissus sp. aff. pseudonarcissus, Galanthus sp., and Galanthus elwesii we have identified a para-para' C-C phenol coupling cytochrome P450, CYP96T1, capable of forming the products (10bR,4aS)-noroxomaritidine and (10bS,4aR)-noroxomaritidine from 4'-O-methylnorbelladine. CYP96T1 was also shown to catalyzed formation of the para-ortho' phenol coupled product, N-demethylnarwedine, as less than 1% of the total product. CYP96T1 co-expresses with the previously characterized norbelladine 4'-O-methyltransferase. The discovery of CYP96T1 is of special interest because it catalyzes the first major branch in Amaryllidaceae alkaloid biosynthesis. CYP96T1 is also the first phenol-coupling enzyme characterized from a monocot.
Keywords: Amaryllidaceae alkaloids; cytochrome P450; phenol coupling; secondary metabolism; transcriptomics.
Figures





References
-
- Barton D. H. R., Kirby G. W., Taylor J. B., Thomas G. M. (1961). The biosynthesis of Amaryllidaceae alkaloids. Proc. Chem. Soc. 254–255. - PubMed
-
- Barton D. H. R., Kirby G. W., Thomas G. M. (1963). Phenol oxidation and biosynthesis. Part VI. The biogenesis of Amaryllidaceae alkaloids. J. Chem. Soc. 1963, 4545–4558. 10.1039/jr9630004545 - DOI
-
- Battersby A. R., Bink R., Breuer S. W. (1961a). Biosynthesis in the Amaryllidaceae: incorporation of norbelladine into lycorine and norpluvine. Proc. Chem. Soc. 243.
-
- Battersby A. R., Binks R. (1960). Biosynthesis of lycorine. Proc. Chem. Soc. 410–411. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

