The BAF chromatin remodelling complex is an epigenetic regulator of lineage specification in the early mouse embryo
- PMID: 26952987
- PMCID: PMC4852518
- DOI: 10.1242/dev.131961
The BAF chromatin remodelling complex is an epigenetic regulator of lineage specification in the early mouse embryo
Abstract
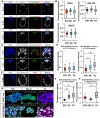
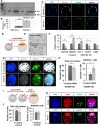
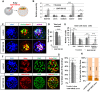
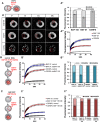
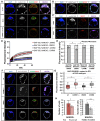
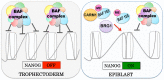
Dynamic control of gene expression is essential for the development of a totipotent zygote into an embryo with defined cell lineages. The accessibility of genes responsible for cell specification to transcriptional machinery is dependent on chromatin remodelling complexes such as the SWI\SNF (BAF) complex. However, the role of the BAF complex in early mouse development has remained unclear. Here, we demonstrate that BAF155, a major BAF complex subunit, regulates the assembly of the BAF complex in vivo and regulates lineage specification of the mouse blastocyst. We find that associations of BAF155 with other BAF complex subunits become enriched in extra-embryonic lineages just prior to implantation. This enrichment is attributed to decreased mobility of BAF155 in extra-embryonic compared with embryonic lineages. Downregulation of BAF155 leads to increased expression of the pluripotency marker Nanog and its ectopic expression in extra-embryonic lineages, whereas upregulation of BAF155 leads to the upregulation of differentiation markers. Finally, we show that the arginine methyltransferase CARM1 methylates BAF155, which differentially influences assembly of the BAF complex between the lineages and the expression of pluripotency markers. Together, our results indicate a novel role of BAF-dependent chromatin remodelling in mouse development via regulation of lineage specification.
Keywords: BAF complex; Chromatin remodelling; Epigenetics; Lineage specification; Mouse embryo; Pluripotency; SMARCC1.
© 2016. Published by The Company of Biologists Ltd.
Figures


Similar articles
-
Epigenetic regulation by BAF (mSWI/SNF) chromatin remodeling complexes is indispensable for embryonic development.Cell Cycle. 2016 May 18;15(10):1317-24. doi: 10.1080/15384101.2016.1160984. Epub 2016 Mar 17. Cell Cycle. 2016. PMID: 26986003 Free PMC article.
-
Requisite Chromatin Remodeling for Myeloid and Erythroid Lineage Differentiation from Erythromyeloid Progenitors.Cell Rep. 2020 Nov 17;33(7):108395. doi: 10.1016/j.celrep.2020.108395. Cell Rep. 2020. PMID: 33207205 Free PMC article.
-
RBM15 Modulates the Function of Chromatin Remodeling Factor BAF155 Through RNA Methylation in Developing Cortex.Mol Neurobiol. 2019 Nov;56(11):7305-7320. doi: 10.1007/s12035-019-1595-1. Epub 2019 Apr 24. Mol Neurobiol. 2019. PMID: 31020615
-
Epigenetic control of cell fate in mouse blastocysts: the role of covalent histone modifications and chromatin remodeling.Mol Reprod Dev. 2014 Feb;81(2):171-82. doi: 10.1002/mrd.22219. Epub 2013 Aug 13. Mol Reprod Dev. 2014. PMID: 23893501 Free PMC article. Review.
-
Epigenetic reorganization during early embryonic lineage specification.Genes Genomics. 2022 Mar;44(3):379-387. doi: 10.1007/s13258-021-01213-w. Epub 2022 Feb 8. Genes Genomics. 2022. PMID: 35133623 Review.
Cited by
-
Zebrafish smarcc1a mutants reveal requirements for BAF chromatin remodeling complexes in distinguishing the atrioventricular canal from the cardiac chambers.Dev Dyn. 2024 Jan;253(1):157-172. doi: 10.1002/dvdy.595. Epub 2023 May 3. Dev Dyn. 2024. PMID: 37083132 Free PMC article.
-
Intrinsic Disorder of the BAF Complex: Roles in Chromatin Remodeling and Disease Development.Int J Mol Sci. 2019 Oct 23;20(21):5260. doi: 10.3390/ijms20215260. Int J Mol Sci. 2019. PMID: 31652801 Free PMC article.
-
CARM1 and Paraspeckles Regulate Pre-implantation Mouse Embryo Development.Cell. 2018 Dec 13;175(7):1902-1916.e13. doi: 10.1016/j.cell.2018.11.027. Cell. 2018. PMID: 30550788 Free PMC article.
-
Principles of Self-Organization of the Mammalian Embryo.Cell. 2020 Dec 10;183(6):1467-1478. doi: 10.1016/j.cell.2020.11.003. Cell. 2020. PMID: 33306953 Free PMC article. Review.
-
A monoastral mitotic spindle determines lineage fate and position in the mouse embryo.Nat Cell Biol. 2022 Feb;24(2):155-167. doi: 10.1038/s41556-021-00826-3. Epub 2022 Jan 31. Nat Cell Biol. 2022. PMID: 35102267 Free PMC article.
References
-
- Bischoff M., Parfitt D.-E. and Zernicka-Goetz M. (2008). Formation of the embryonic-abembryonic axis of the mouse blastocyst: relationships between orientation of early cleavage divisions and pattern of symmetric/asymmetric divisions. Development 135, 953-962. 10.1242/dev.014316 - DOI - PMC - PubMed
-
- Bultman S., Gebuhr T., Yee D., La Mantia C., Nicholson J., Gilliam A., Randazzo F., Metzger D., Chambon P., Crabtree G. et al. (2000). A Brg1 null mutation in the mouse reveals functional differences among mammalian SWI/SNF complexes. Mol. Cell 6, 1287-1295. 10.1016/S1097-2765(00)00127-1 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

