Inositol Hexakisphosphate Kinase 1 (IP6K1) Regulates Inositol Synthesis in Mammalian Cells
- PMID: 26953345
- PMCID: PMC4865894
- DOI: 10.1074/jbc.M116.714816
Inositol Hexakisphosphate Kinase 1 (IP6K1) Regulates Inositol Synthesis in Mammalian Cells
Abstract
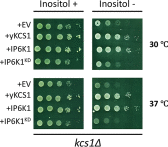
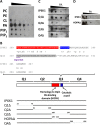
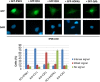
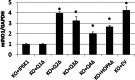
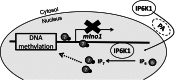
myo-Inositol, the precursor of all inositol compounds, has pivotal roles in cell metabolism and signaling pathways. Although physiological studies indicate a strong correlation between abnormal intracellular inositol levels and neurological disorders, very little is known about the regulation of inositol synthesis in mammalian cells. In this study, we report that IP6K1, an inositol hexakisphosphate kinase that catalyzes the synthesis of inositol pyrophosphate, regulates inositol synthesis in mammalian cells. Ip6k1 ablation led to profound changes in DNA methylation and expression of Isyna1 (designated mIno1), which encodes the rate-limiting enzyme inositol-3-phosphate synthase. Interestingly, IP6K1 preferentially bound to the phospholipid phosphatidic acid, and this binding was required for IP6K1 nuclear localization and the regulation of mIno1 transcription. This is the first demonstration of IP6K1 as a novel negative regulator of inositol synthesis in mammalian cells.
Keywords: ISYNA1; Opi1; genetics; inositol; inositol phosphate; inositol pyrophosphate; lipid; metabolism; phosphatidic acid.
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures


Similar articles
-
Phosphatidic acid inhibits inositol synthesis by inducing nuclear translocation of kinase IP6K1 and repression of myo-inositol-3-P synthase.J Biol Chem. 2022 Sep;298(9):102363. doi: 10.1016/j.jbc.2022.102363. Epub 2022 Aug 10. J Biol Chem. 2022. PMID: 35963434 Free PMC article.
-
Inositol hexakisphosphate kinase-1 interacts with perilipin1 to modulate lipolysis.Int J Biochem Cell Biol. 2016 Sep;78:149-155. doi: 10.1016/j.biocel.2016.06.018. Epub 2016 Jun 29. Int J Biochem Cell Biol. 2016. PMID: 27373682 Free PMC article.
-
Deletion of inositol hexakisphosphate kinase 1 (IP6K1) reduces cell migration and invasion, conferring protection from aerodigestive tract carcinoma in mice.Cell Signal. 2016 Aug;28(8):1124-36. doi: 10.1016/j.cellsig.2016.04.011. Epub 2016 Apr 30. Cell Signal. 2016. PMID: 27140681 Free PMC article.
-
Insights into the roles of inositol hexakisphosphate kinase 1 (IP6K1) in mammalian cellular processes.J Biol Chem. 2024 Apr;300(4):107116. doi: 10.1016/j.jbc.2024.107116. Epub 2024 Feb 24. J Biol Chem. 2024. PMID: 38403246 Free PMC article. Review.
-
Targeting the Inositol Pyrophosphate Biosynthetic Enzymes in Metabolic Diseases.Molecules. 2020 Mar 19;25(6):1403. doi: 10.3390/molecules25061403. Molecules. 2020. PMID: 32204420 Free PMC article. Review.
Cited by
-
Inositols in Insulin Signaling and Glucose Metabolism.Int J Endocrinol. 2018 Nov 25;2018:1968450. doi: 10.1155/2018/1968450. eCollection 2018. Int J Endocrinol. 2018. PMID: 30595691 Free PMC article. Review.
-
Target of Rapamycin Complex 1 (TORC1), Protein Kinase A (PKA) and Cytosolic pH Regulate a Transcriptional Circuit for Lipid Droplet Formation.Int J Mol Sci. 2021 Aug 20;22(16):9017. doi: 10.3390/ijms22169017. Int J Mol Sci. 2021. PMID: 34445723 Free PMC article.
-
The Key Role of IP6K: A Novel Target for Anticancer Treatments?Molecules. 2020 Sep 25;25(19):4401. doi: 10.3390/molecules25194401. Molecules. 2020. PMID: 32992691 Free PMC article. Review.
-
Inositol depletion regulates phospholipid metabolism and activates stress signaling in HEK293T cells.Biochim Biophys Acta Mol Cell Biol Lipids. 2022 Jun;1867(6):159137. doi: 10.1016/j.bbalip.2022.159137. Epub 2022 Mar 3. Biochim Biophys Acta Mol Cell Biol Lipids. 2022. PMID: 35247568 Free PMC article.
-
Valproate regulates inositol synthesis by reducing expression of myo-inositol-3-phosphate synthase.Sci Rep. 2023 Sep 8;13(1):14844. doi: 10.1038/s41598-023-41936-2. Sci Rep. 2023. PMID: 37684289 Free PMC article.
References
-
- Streb H., Irvine R. F., Berridge M. J., and Schulz I. (1983) Release of Ca2+ from a nonmitochondrial intracellular store in pancreatic acinar cells by inositol-1,4,5-trisphosphate. Nature 306, 67–69 - PubMed
-
- Shi Y., Azab A. N., Thompson M. N., and Greenberg M. L. (2006) Inositol phosphates and phosphoinositides in health and disease. Subcell. Biochem. 39, 265–292 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

