ARHGEF17 is an essential spindle assembly checkpoint factor that targets Mps1 to kinetochores
- PMID: 26953350
- PMCID: PMC4792069
- DOI: 10.1083/jcb.201408089
ARHGEF17 is an essential spindle assembly checkpoint factor that targets Mps1 to kinetochores
Abstract
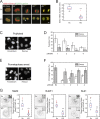
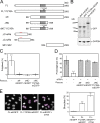
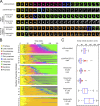
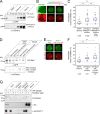
To prevent genome instability, mitotic exit is delayed until all chromosomes are properly attached to the mitotic spindle by the spindle assembly checkpoint (SAC). In this study, we characterized the function of ARHGEF17, identified in a genome-wide RNA interference screen for human mitosis genes. Through a series of quantitative imaging, biochemical, and biophysical experiments, we showed that ARHGEF17 is essential for SAC activity, because it is the major targeting factor that controls localization of the checkpoint kinase Mps1 to the kinetochore. This mitotic function is mediated by direct interaction of the central domain of ARHGEF17 with Mps1, which is autoregulated by the activity of Mps1 kinase, for which ARHGEF17 is a substrate. This mitosis-specific role is independent of ARHGEF17's RhoGEF activity in interphase. Our study thus assigns a new mitotic function to ARHGEF17 and reveals the molecular mechanism for a key step in SAC establishment.
© 2016 Isokane et al.
Figures




Comment in
-
ARHGEF17 sets the timer for retention of Mps1 at kinetochores.J Cell Biol. 2016 Mar 14;212(6):615-6. doi: 10.1083/jcb.201602060. Epub 2016 Mar 7. J Cell Biol. 2016. PMID: 26953355 Free PMC article.
Similar articles
-
ARHGEF17 sets the timer for retention of Mps1 at kinetochores.J Cell Biol. 2016 Mar 14;212(6):615-6. doi: 10.1083/jcb.201602060. Epub 2016 Mar 7. J Cell Biol. 2016. PMID: 26953355 Free PMC article.
-
Interactions between N-terminal Modules in MPS1 Enable Spindle Checkpoint Silencing.Cell Rep. 2019 Feb 19;26(8):2101-2112.e6. doi: 10.1016/j.celrep.2019.01.017. Cell Rep. 2019. PMID: 30784592
-
CELL DIVISION CYCLE. Competition between MPS1 and microtubules at kinetochores regulates spindle checkpoint signaling.Science. 2015 Jun 12;348(6240):1264-7. doi: 10.1126/science.aaa4055. Epub 2015 Jun 11. Science. 2015. PMID: 26068855
-
Orchestration of the spindle assembly checkpoint by CDK1-cyclin B1.FEBS Lett. 2019 Oct;593(20):2889-2907. doi: 10.1002/1873-3468.13591. Epub 2019 Sep 13. FEBS Lett. 2019. PMID: 31469407 Review.
-
Mps1 kinase functions in mitotic spindle assembly and error correction.Trends Biochem Sci. 2025 May;50(5):438-453. doi: 10.1016/j.tibs.2025.02.005. Epub 2025 Mar 12. Trends Biochem Sci. 2025. PMID: 40082122 Review.
Cited by
-
A commercial ARHGEF17/TEM4 antibody cross-reacts with Nuclear Mitotic Apparatus protein 1 (NuMA).PLoS One. 2022 Jul 1;17(7):e0268848. doi: 10.1371/journal.pone.0268848. eCollection 2022. PLoS One. 2022. PMID: 35776709 Free PMC article.
-
Inward Outward Signaling in Ovarian Cancer: Morpho-Phospho-Proteomic Profiling Upon Application of Hypoxia and Shear Stress Characterizes the Adaptive Plasticity of OVCAR-3 and SKOV-3 Cells.Front Oncol. 2022 Feb 14;11:746411. doi: 10.3389/fonc.2021.746411. eCollection 2021. Front Oncol. 2022. PMID: 35251951 Free PMC article.
-
Game-theoretic link relevance indexing on genome-wide expression dataset identifies putative salient genes with potential etiological and diapeutics role in colorectal cancer.Sci Rep. 2022 Aug 4;12(1):13409. doi: 10.1038/s41598-022-17266-0. Sci Rep. 2022. PMID: 35927308 Free PMC article.
-
Experimental and computational framework for a dynamic protein atlas of human cell division.Nature. 2018 Sep;561(7723):411-415. doi: 10.1038/s41586-018-0518-z. Epub 2018 Sep 10. Nature. 2018. PMID: 30202089 Free PMC article.
-
Case report: Expanding the phenotype of ARHGEF17 mutations from increased intracranial aneurysm risk to a neurodevelopmental disease.Front Neurol. 2022 Oct 20;13:1017654. doi: 10.3389/fneur.2022.1017654. eCollection 2022. Front Neurol. 2022. PMID: 36341116 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

