The Autophagy-Lysosomal Pathway in Neurodegeneration: A TFEB Perspective
- PMID: 26968346
- PMCID: PMC4928589
- DOI: 10.1016/j.tins.2016.02.002
The Autophagy-Lysosomal Pathway in Neurodegeneration: A TFEB Perspective
Abstract
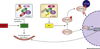
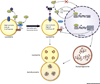
The autophagy-lysosomal pathway (ALP) is involved in the degradation of long-lived proteins. Deficits in the ALP result in protein aggregation, the generation of toxic protein species, and accumulation of dysfunctional organelles, which are hallmarks of Alzheimer's disease (AD), Parkinson's disease (PD), Huntington's disease (HD), and prion disease. Decades of research have therefore focused on enhancing the ALP in neurodegenerative diseases. More recently, transcription factor EB (TFEB), a major regulator of autophagy and lysosomal biogenesis, has emerged as a leading factor in addressing disease pathology. We review the regulation of the ALP and TFEB and their impact on neurodegenerative diseases. We also offer our perspective on the complex role of autophagy and TFEB in disease pathogenesis and its therapeutic implications through the examination of prion disease.
Keywords: TFEB; autophagy–lysosomal pathway; neurodegenerative disease; prion disease.
Copyright © 2016. Published by Elsevier Ltd.
Figures

References
-
- Sardiello M, et al. A gene network regulating lysosomal biogenesis and function. Science. 2009;325:473–477. - PubMed
-
- Komatsu M, et al. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature. 2006;441:880–884. - PubMed
-
- Hara T, et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature. 2006;441:885–889. - PubMed
-
- Menzies FM, et al. Compromised autophagy and neuro-degenerative diseases. Nat. Rev. Neurosci. 2015;16:345–357. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

