Biofilms on Hospital Shower Hoses: Characterization and Implications for Nosocomial Infections
- PMID: 26969701
- PMCID: PMC4836434
- DOI: 10.1128/AEM.03529-15
Biofilms on Hospital Shower Hoses: Characterization and Implications for Nosocomial Infections
Abstract
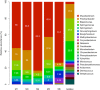
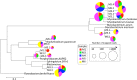
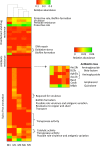
Although the source of drinking water (DW) used in hospitals is commonly disinfected, biofilms forming on water pipelines are a refuge for bacteria, including possible pathogens that survive different disinfection strategies. These biofilm communities are only beginning to be explored by culture-independent techniques that circumvent the limitations of conventional monitoring efforts. Hence, theories regarding the frequency of opportunistic pathogens in DW biofilms and how biofilm members withstand high doses of disinfectants and/or chlorine residuals in the water supply remain speculative. The aim of this study was to characterize the composition of microbial communities growing on five hospital shower hoses using both 16S rRNA gene sequencing of bacterial isolates and whole-genome shotgun metagenome sequencing. The resulting data revealed a Mycobacterium-like population, closely related to Mycobacterium rhodesiae and Mycobacterium tusciae, to be the predominant taxon in all five samples, and its nearly complete draft genome sequence was recovered. In contrast, the fraction recovered by culture was mostly affiliated with Proteobacteria, including members of the genera Sphingomonas, Blastomonas, and Porphyrobacter.The biofilm community harbored genes related to disinfectant tolerance (2.34% of the total annotated proteins) and a lower abundance of virulence determinants related to colonization and evasion of the host immune system. Additionally, genes potentially conferring resistance to β-lactam, aminoglycoside, amphenicol, and quinolone antibiotics were detected. Collectively, our results underscore the need to understand the microbiome of DW biofilms using metagenomic approaches. This information might lead to more robust management practices that minimize the risks associated with exposure to opportunistic pathogens in hospitals.
Copyright © 2016, American Society for Microbiology. All Rights Reserved.
Figures
References
-
- Liu R, Yu Z, Guo H, Liu M, Zhang H, Yang M. 2012. Pyrosequencing analysis of eukaryotic and bacterial communities in faucet biofilms. Sci Total Environ 435–436:124–131. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

