Sp5 and Sp8 recruit β-catenin and Tcf1-Lef1 to select enhancers to activate Wnt target gene transcription
- PMID: 26969725
- PMCID: PMC4822596
- DOI: 10.1073/pnas.1519994113
Sp5 and Sp8 recruit β-catenin and Tcf1-Lef1 to select enhancers to activate Wnt target gene transcription
Abstract
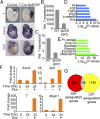
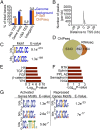
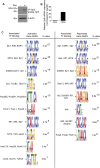
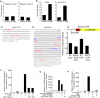
The ancient, highly conserved, Wnt signaling pathway regulates cell fate in all metazoans. We have previously shown that combined null mutations of the specificity protein (Sp) 1/Klf-like zinc-finger transcription factors Sp5 and Sp8 (i.e., Sp5/8) result in an embryonic phenotype identical to that observed when core components of the Wnt/β-catenin pathway are mutated; however, their role in Wnt signal transduction is unknown. Here, we show in mouse embryos and differentiating embryonic stem cells that Sp5/8 are gene-specific transcriptional coactivators in the Wnt/β-catenin pathway. Sp5/8 bind directly to GC boxes in Wnt target gene enhancers and to adjacent, or distally positioned, chromatin-bound T-cell factor (Tcf) 1/lymphoid enhancer factor (Lef) 1 to facilitate recruitment of β-catenin to target gene enhancers. Because Sp5 is itself directly activated by Wnt signals, we propose that Sp5 is a Wnt/β-catenin pathway-specific transcript on factor that functions in a feed-forward loop to robustly activate select Wnt target genes.
Keywords: Sp5/Sp8; Tcf/Lef; Wnt; stem cells; transcription.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


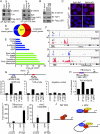

Similar articles
-
β-Catenin-independent activation of TCF1/LEF1 in human hematopoietic tumor cells through interaction with ATF2 transcription factors.PLoS Genet. 2013;9(8):e1003603. doi: 10.1371/journal.pgen.1003603. Epub 2013 Aug 15. PLoS Genet. 2013. PMID: 23966864 Free PMC article.
-
Zfp703 Is a Wnt/β-Catenin Feedback Suppressor Targeting the β-Catenin/Tcf1 Complex.Mol Cell Biol. 2016 May 31;36(12):1793-802. doi: 10.1128/MCB.01010-15. Print 2016 Jun 15. Mol Cell Biol. 2016. PMID: 27090637 Free PMC article.
-
Function of Wnt/β-catenin in counteracting Tcf3 repression through the Tcf3-β-catenin interaction.Development. 2012 Jun;139(12):2118-29. doi: 10.1242/dev.076067. Epub 2012 May 9. Development. 2012. PMID: 22573616 Free PMC article.
-
Wnt/β-Catenin Signaling Pathway Governs a Full Program for Dopaminergic Neuron Survival, Neurorescue and Regeneration in the MPTP Mouse Model of Parkinson's Disease.Int J Mol Sci. 2018 Nov 24;19(12):3743. doi: 10.3390/ijms19123743. Int J Mol Sci. 2018. PMID: 30477246 Free PMC article. Review.
-
A WNTer revisit: new faces of β-catenin and TCFs in pluripotency.Sci Signal. 2011 Sep 27;4(193):pe41. doi: 10.1126/scisignal.2002436. Sci Signal. 2011. PMID: 21971038 Review.
Cited by
-
Transcription factor specificity protein (SP) family in renal physiology and diseases.PeerJ. 2025 Jan 20;13:e18820. doi: 10.7717/peerj.18820. eCollection 2025. PeerJ. 2025. PMID: 39850832 Free PMC article. Review.
-
G9a regulates tumorigenicity and stemness through genome-wide DNA methylation reprogramming in non-small cell lung cancer.Clin Epigenetics. 2020 Jun 17;12(1):88. doi: 10.1186/s13148-020-00879-5. Clin Epigenetics. 2020. PMID: 32552834 Free PMC article.
-
FOXO1-regulated lncRNA LINC01197 inhibits pancreatic adenocarcinoma cell proliferation by restraining Wnt/β-catenin signaling.J Exp Clin Cancer Res. 2019 Apr 26;38(1):179. doi: 10.1186/s13046-019-1174-3. J Exp Clin Cancer Res. 2019. PMID: 31027497 Free PMC article.
-
Wnt target enhancer regulation by a CDX/TCF transcription factor collective and a novel DNA motif.Nucleic Acids Res. 2021 Sep 7;49(15):8625-8641. doi: 10.1093/nar/gkab657. Nucleic Acids Res. 2021. PMID: 34358319 Free PMC article.
-
Wnt target genes and where to find them.F1000Res. 2017 May 24;6:746. doi: 10.12688/f1000research.11034.1. eCollection 2017. F1000Res. 2017. PMID: 28649368 Free PMC article. Review.
References
-
- Clevers H, Nusse R. Wnt/β-catenin signaling and disease. Cell. 2012;149(6):1192–1205. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

