Repurposing the CRISPR-Cas9 system for targeted DNA methylation
- PMID: 26969735
- PMCID: PMC4937303
- DOI: 10.1093/nar/gkw159
Repurposing the CRISPR-Cas9 system for targeted DNA methylation
Abstract
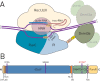
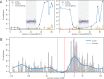
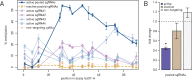
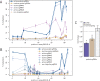
Epigenetic studies relied so far on correlations between epigenetic marks and gene expression pattern. Technologies developed for epigenome editing now enable direct study of functional relevance of precise epigenetic modifications and gene regulation. The reversible nature of epigenetic modifications, including DNA methylation, has been already exploited in cancer therapy for remodeling the aberrant epigenetic landscape. However, this was achieved non-selectively using epigenetic inhibitors. Epigenetic editing at specific loci represents a novel approach that might selectively and heritably alter gene expression. Here, we developed a CRISPR-Cas9-based tool for specific DNA methylation consisting of deactivated Cas9 (dCas9) nuclease and catalytic domain of the DNA methyltransferase DNMT3A targeted by co-expression of a guide RNA to any 20 bp DNA sequence followed by the NGG trinucleotide. We demonstrated targeted CpG methylation in a ∼35 bp wide region by the fusion protein. We also showed that multiple guide RNAs could target the dCas9-DNMT3A construct to multiple adjacent sites, which enabled methylation of a larger part of the promoter. DNA methylation activity was specific for the targeted region and heritable across mitotic divisions. Finally, we demonstrated that directed DNA methylation of a wider promoter region of the target loci IL6ST and BACH2 decreased their expression.
© The Author(s) 2016. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


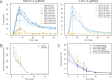
References
-
- Urnov F.D., Rebar E.J., Holmes M.C., Zhang H.S., Gregory P.D. Genome editing with engineered zinc finger nucleases. Nat. Rev. Genet. 2010;11:636–646. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

