Subcellular Compartmentalization and Trafficking of the Biosynthetic Machinery for Fungal Melanin
- PMID: 26972005
- PMCID: PMC4805463
- DOI: 10.1016/j.celrep.2016.02.059
Subcellular Compartmentalization and Trafficking of the Biosynthetic Machinery for Fungal Melanin
Abstract
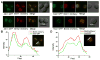
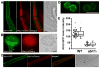
Protection by melanin depends on its subcellular location. Although most filamentous fungi synthesize melanin via a polyketide synthase pathway, where and how melanin biosynthesis occurs and how it is deposited as extracellular granules remain elusive. Using a forward genetic screen in the pathogen Aspergillus fumigatus, we find that mutations in an endosomal sorting nexin abolish melanin cell-wall deposition. We find that all enzymes involved in the early steps of melanin biosynthesis are recruited to endosomes through a non-conventional secretory pathway. In contrast, late melanin enzymes accumulate in the cell wall. Such subcellular compartmentalization of the melanin biosynthetic machinery occurs in both A. fumigatus and A. nidulans. Thus, fungal melanin biosynthesis appears to be initiated in endosomes with exocytosis leading to melanin extracellular deposition, much like the synthesis and trafficking of mammalian melanin in endosomally derived melanosomes.
Copyright © 2016 The Authors. Published by Elsevier Inc. All rights reserved.
Figures


Similar articles
-
Compartmentalization of Melanin Biosynthetic Enzymes Contributes to Self-Defense against Intermediate Compound Scytalone in Botrytis cinerea.mBio. 2021 Mar 23;12(2):e00007-21. doi: 10.1128/mBio.00007-21. mBio. 2021. PMID: 33758088 Free PMC article.
-
Targeting of Specialized Metabolites Biosynthetic Enzymes to Membranes and Vesicles by Posttranslational Palmitoylation: A Mechanism of Non-Conventional Traffic and Secretion of Fungal Metabolites.Int J Mol Sci. 2024 Jan 19;25(2):1224. doi: 10.3390/ijms25021224. Int J Mol Sci. 2024. PMID: 38279221 Free PMC article. Review.
-
Interactions between Melanin Enzymes and Their Atypical Recruitment to the Secretory Pathway by Palmitoylation.mBio. 2016 Nov 22;7(6):e01925-16. doi: 10.1128/mBio.01925-16. mBio. 2016. PMID: 27879337 Free PMC article.
-
SNX-BAR-mediated endosome tubulation is co-ordinated with endosome maturation.Traffic. 2012 Jan;13(1):94-107. doi: 10.1111/j.1600-0854.2011.01297.x. Epub 2011 Oct 31. Traffic. 2012. PMID: 21973056
-
The fungal RABOME: RAB GTPases acting in the endocytic and exocytic pathways of Aspergillus nidulans (with excursions to other filamentous fungi).Mol Microbiol. 2021 Jul;116(1):53-70. doi: 10.1111/mmi.14716. Epub 2021 Apr 3. Mol Microbiol. 2021. PMID: 33724562 Review.
Cited by
-
Compartmentalization of Melanin Biosynthetic Enzymes Contributes to Self-Defense against Intermediate Compound Scytalone in Botrytis cinerea.mBio. 2021 Mar 23;12(2):e00007-21. doi: 10.1128/mBio.00007-21. mBio. 2021. PMID: 33758088 Free PMC article.
-
The Genetics and Biochemistry of Cell Wall Structure and Synthesis in Neurospora crassa, a Model Filamentous Fungus.Front Microbiol. 2019 Oct 10;10:2294. doi: 10.3389/fmicb.2019.02294. eCollection 2019. Front Microbiol. 2019. PMID: 31649638 Free PMC article. Review.
-
The Transcriptomic and Phenotypic Response of the Melanized Yeast Exophiala dermatitidis to Ionizing Particle Exposure.Front Microbiol. 2021 Jan 12;11:609996. doi: 10.3389/fmicb.2020.609996. eCollection 2020. Front Microbiol. 2021. PMID: 33510728 Free PMC article.
-
The spatial organization of sphingofungin biosynthesis in Aspergillus fumigatus and its cross-interaction with sphingolipid metabolism.mBio. 2024 Mar 13;15(3):e0019524. doi: 10.1128/mbio.00195-24. Epub 2024 Feb 21. mBio. 2024. PMID: 38380921 Free PMC article.
-
Targeting of Specialized Metabolites Biosynthetic Enzymes to Membranes and Vesicles by Posttranslational Palmitoylation: A Mechanism of Non-Conventional Traffic and Secretion of Fungal Metabolites.Int J Mol Sci. 2024 Jan 19;25(2):1224. doi: 10.3390/ijms25021224. Int J Mol Sci. 2024. PMID: 38279221 Free PMC article. Review.
References
-
- Alviano CS, Farbiarz SR, De Souza W, Angluster J, Travassos LR. Characterization of Fonsecaea pedrosoi melanin. J Gen Microbiol. 1991;137:837–844. - PubMed
-
- Banci L, Bertini I, McGreevy KS, Rosato A. Molecular recognition in copper trafficking. Natural product reports. 2010;27:695–710. - PubMed
-
- Barral DC, Seabra MC. The melanosome as a model to study organelle motility in mammals. Pigment cell research / sponsored by the European Society for Pigment Cell Research and the International Pigment Cell Society. 2004;17:111–118. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

