Gram-negative and -positive bacteria differentiation in blood culture samples by headspace volatile compound analysis
- PMID: 26973820
- PMCID: PMC4788920
- DOI: 10.1186/s40709-016-0040-0
Gram-negative and -positive bacteria differentiation in blood culture samples by headspace volatile compound analysis
Abstract
Background: Identification of microorganisms in positive blood cultures still relies on standard techniques such as Gram staining followed by culturing with definite microorganism identification. Alternatively, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry or the analysis of headspace volatile compound (VC) composition produced by cultures can help to differentiate between microorganisms under experimental conditions. This study assessed the efficacy of volatile compound based microorganism differentiation into Gram-negatives and -positives in unselected positive blood culture samples from patients.
Methods: Headspace gas samples of positive blood culture samples were transferred to sterilized, sealed, and evacuated 20 ml glass vials and stored at -30 °C until batch analysis. Headspace gas VC content analysis was carried out via an auto sampler connected to an ion-molecule reaction mass spectrometer (IMR-MS). Measurements covered a mass range from 16 to 135 u including CO2, H2, N2, and O2. Prediction rules for microorganism identification based on VC composition were derived using a training data set and evaluated using a validation data set within a random split validation procedure.
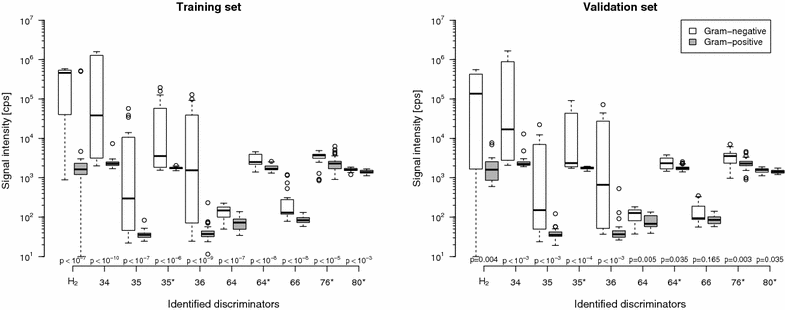
Results: One-hundred-fifty-two aerobic samples growing 27 Gram-negatives, 106 Gram-positives, and 19 fungi and 130 anaerobic samples growing 37 Gram-negatives, 91 Gram-positives, and two fungi were analysed. In anaerobic samples, ten discriminators were identified by the random forest method allowing for bacteria differentiation into Gram-negative and -positive (error rate: 16.7 % in validation data set). For aerobic samples the error rate was not better than random.
Conclusions: In anaerobic blood culture samples of patients IMR-MS based headspace VC composition analysis facilitates bacteria differentiation into Gram-negative and -positive.
Keywords: Blood culture; Chemical ionization; Gram identification; Mass spectrometry; Prediction rule; Volatile compound.
Figures
Similar articles
-
Identification of pathogenic microorganisms directly from positive blood vials by matrix-assisted laser desorption/ionization time of flight mass spectrometry.APMIS. 2013 Sep;121(9):871-7. doi: 10.1111/apm.12050. Epub 2013 Jan 18. APMIS. 2013. PMID: 23331371
-
Volatile organic compound analysis by ion molecule reaction mass spectrometry for Gram-positive bacteria differentiation.Eur J Clin Microbiol Infect Dis. 2012 Nov;31(11):3007-13. doi: 10.1007/s10096-012-1654-2. Epub 2012 Jul 11. Eur J Clin Microbiol Infect Dis. 2012. PMID: 22782437
-
Volatile compound profiling for the identification of Gram-negative bacteria by ion-molecule reaction-mass spectrometry.J Appl Microbiol. 2012 Nov;113(5):1097-105. doi: 10.1111/j.1365-2672.2012.05414.x. Epub 2012 Aug 21. J Appl Microbiol. 2012. PMID: 22830412
-
Microorganisms direct identification from blood culture by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry.Clin Microbiol Infect. 2011 Apr;17(4):546-51. doi: 10.1111/j.1469-0691.2010.03257.x. Clin Microbiol Infect. 2011. PMID: 20456452
-
Discrimination of bacteria by rapid sensing their metabolic volatiles using an aspiration-type ion mobility spectrometer (a-IMS) and gas chromatography-mass spectrometry GC-MS.Anal Chim Acta. 2017 Aug 22;982:209-217. doi: 10.1016/j.aca.2017.06.031. Epub 2017 Jun 22. Anal Chim Acta. 2017. PMID: 28734362
Cited by
-
Comprehensive volatile metabolic fingerprinting of bacterial and fungal pathogen groups.J Breath Res. 2018 Jan 3;12(2):026001. doi: 10.1088/1752-7163/aa8f7f. J Breath Res. 2018. PMID: 28952968 Free PMC article.
-
Machine learning for the meta-analyses of microbial pathogens' volatile signatures.Sci Rep. 2018 Feb 20;8(1):3360. doi: 10.1038/s41598-018-21544-1. Sci Rep. 2018. PMID: 29463885 Free PMC article.
-
The volatile molecular profiles of seven Streptococcus pneumoniae serotypes.J Chromatogr B Analyt Technol Biomed Life Sci. 2018 Oct 1;1096:208-213. doi: 10.1016/j.jchromb.2018.08.032. Epub 2018 Aug 29. J Chromatogr B Analyt Technol Biomed Life Sci. 2018. PMID: 30179753 Free PMC article.
-
Modern approaches for detection of volatile organic compounds in metabolic studies focusing on pathogenic bacteria: Current state of the art.J Pharm Anal. 2024 Apr;14(4):100898. doi: 10.1016/j.jpha.2023.11.005. Epub 2023 Nov 28. J Pharm Anal. 2024. PMID: 38634063 Free PMC article. Review.
-
Sniffing Bacteria with a Carbon-Dot Artificial Nose.Nanomicro Lett. 2021 Apr 20;13(1):112. doi: 10.1007/s40820-021-00610-w. Nanomicro Lett. 2021. PMID: 34138310 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

