Engineering Modular Viral Scaffolds for Targeted Bacterial Population Editing
- PMID: 26973885
- PMCID: PMC4785837
- DOI: 10.1016/j.cels.2015.08.013
Engineering Modular Viral Scaffolds for Targeted Bacterial Population Editing
Abstract
Bacteria are central to human health and disease, but existing tools to edit microbial consortia are limited. For example, broad-spectrum antibiotics are unable to accurately manipulate bacterial communities. Bacteriophages can provide highly specific targeting of bacteria, but assembling well-defined phage cocktails solely with natural phages can be a time-, labor- and cost-intensive process. Here, we present a synthetic-biology strategy to modulate phage host ranges by engineering phage genomes in Saccharomyces cerevisiae. We used this technology to redirect Escherichia coli phage scaffolds to target pathogenic Yersinia and Klebsiella bacteria, and conversely, Klebsiella phage scaffolds to target E. coli by modular swapping of phage tail components. The synthetic phages achieved efficient killing of their new target bacteria and were used to selectively remove bacteria from multi-species bacterial communities with cocktails based on common viral scaffolds. We envision that this approach will accelerate phage-biology studies and enable new technologies for bacterial population editing.
Figures





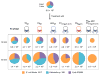
References
-
- Bessler W, Freund-Molbert E, Knufermann H, Rduolph C, Thurow H, Stirm S. A bacteriophage-induced depolymerase active on Klebsiella K11 capsular polysaccharide. Virology. 1973;56:134–151. - PubMed
-
- Brussow H. What is needed for phage therapy to become a reality in Western medicine? Virology. 2012;434:138–142. - PubMed
-
- Carlton RM. Phage therapy: past history and future prospects. Archivum immunologiae et therapiae experimentalis. 1999;47:267–274. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

