RNA-binding protein IGF2BP3 targeting of oncogenic transcripts promotes hematopoietic progenitor proliferation
- PMID: 26974154
- PMCID: PMC4811152
- DOI: 10.1172/JCI80046
RNA-binding protein IGF2BP3 targeting of oncogenic transcripts promotes hematopoietic progenitor proliferation
Abstract
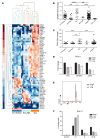
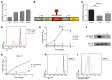
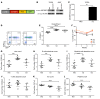
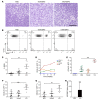
Posttranscriptional control of gene expression is important for defining both normal and pathological cellular phenotypes. In vitro, RNA-binding proteins (RBPs) have recently been shown to play important roles in posttranscriptional regulation; however, the contribution of RBPs to cell specification is not well understood. Here, we determined that the RBP insulin-like growth factor 2 mRNA-binding protein 3 (IGF2BP3) is specifically overexpressed in mixed lineage leukemia-rearranged (MLL-rearranged) B-acute lymphoblastic leukemia (B-ALL), which constitutes a subtype of this malignancy associated with poor prognosis and high risk of relapse. IGF2BP3 was required for the survival of B-ALL cell lines, as knockdown led to decreased proliferation and increased apoptosis. Enforced expression of IGF2BP3 provided murine BM cells with a strong survival advantage, led to proliferation of hematopoietic stem and progenitor cells, and skewed hematopoietic development to the B cell/myeloid lineage. Cross-link immunoprecipitation and high throughput sequencing uncovered the IGF2BP3-regulated transcriptome, which includes oncogenes MYC and CDK6 as direct targets. IGF2BP3 regulated transcripts via targeting elements within 3' untranslated regions (3'UTR), and enforced IGF2BP3 expression in mice resulted in enhanced expression of Myc and Cdk6 in BM. Together, our data suggest that IGF2BP3-mediated targeting of oncogenic transcripts may represent a critical pathogenetic mechanism in MLL-rearranged B-ALL and support IGF2BP3 and its cognate RNA-binding partners as potential therapeutic targets in this disease.
Figures



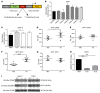

Similar articles
-
Stat5-deficient hematopoiesis is permissive for Myc-induced B-cell leukemogenesis.Oncotarget. 2015 Oct 6;6(30):28961-72. doi: 10.18632/oncotarget.5009. Oncotarget. 2015. PMID: 26338970 Free PMC article.
-
The RNA-binding protein IGF2BP3 is critical for MLL-AF4-mediated leukemogenesis.Leukemia. 2022 Jan;36(1):68-79. doi: 10.1038/s41375-021-01346-7. Epub 2021 Jul 29. Leukemia. 2022. PMID: 34321607 Free PMC article.
-
MLL fusion-driven activation of CDK6 potentiates proliferation in MLL-rearranged infant ALL.Cell Cycle. 2014;13(5):834-44. doi: 10.4161/cc.27757. Cell Cycle. 2014. PMID: 24736461 Free PMC article.
-
Regulation of the MIR155 host gene in physiological and pathological processes.Gene. 2013 Dec 10;532(1):1-12. doi: 10.1016/j.gene.2012.12.009. Epub 2012 Dec 14. Gene. 2013. PMID: 23246696 Review.
-
Expression of FLT3 receptor and response to FLT3 ligand by leukemic cells.Leukemia. 1996 Apr;10(4):588-99. Leukemia. 1996. PMID: 8618433 Review.
Cited by
-
Ubiquitination and deubiquitination in the regulation of N6-methyladenosine functional molecules.J Mol Med (Berl). 2024 Mar;102(3):337-351. doi: 10.1007/s00109-024-02417-9. Epub 2024 Jan 30. J Mol Med (Berl). 2024. PMID: 38289385 Review.
-
The Role of RNA-Binding Proteins in Hematological Malignancies.Int J Mol Sci. 2022 Aug 23;23(17):9552. doi: 10.3390/ijms23179552. Int J Mol Sci. 2022. PMID: 36076951 Free PMC article. Review.
-
Insulin-Like Growth Factor 2 mRNA Binding Protein 3 Promotes Cell Proliferation of Malignant Mesothelioma Cells by Downregulating p27Kip1.Front Oncol. 2022 Jan 19;11:795467. doi: 10.3389/fonc.2021.795467. eCollection 2021. Front Oncol. 2022. PMID: 35127504 Free PMC article.
-
Mechanistic basis of neonatal heart regeneration revealed by transcriptome and histone modification profiling.Proc Natl Acad Sci U S A. 2019 Sep 10;116(37):18455-18465. doi: 10.1073/pnas.1905824116. Epub 2019 Aug 26. Proc Natl Acad Sci U S A. 2019. PMID: 31451669 Free PMC article.
-
IMP3 Stabilization of WNT5B mRNA Facilitates TAZ Activation in Breast Cancer.Cell Rep. 2018 May 29;23(9):2559-2567. doi: 10.1016/j.celrep.2018.04.113. Cell Rep. 2018. PMID: 29847788 Free PMC article.
References
-
- Mullighan CG, Downing JR. Genome-wide profiling of genetic alterations in acute lymphoblastic leukemia: recent insights and future directions. Leukemia. 2009;23(7):1209–1218. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- K08CA133521/CA/NCI NIH HHS/United States
- P30CA016042/CA/NCI NIH HHS/United States
- R41 HG007336/HG/NHGRI NIH HHS/United States
- K08 CA133521/CA/NCI NIH HHS/United States
- AI-28697/AI/NIAID NIH HHS/United States
- R01GM109146/GM/NIGMS NIH HHS/United States
- R01 CA166450/CA/NCI NIH HHS/United States
- P30 CA016042/CA/NCI NIH HHS/United States
- P30 AI028697/AI/NIAID NIH HHS/United States
- T32 CA009120/CA/NCI NIH HHS/United States
- T32 CA009056/CA/NCI NIH HHS/United States
- R01 GM109146/GM/NIGMS NIH HHS/United States
- T32CA009120/CA/NCI NIH HHS/United States
- R21AG042003/AG/NIA NIH HHS/United States
- T32CA009056/CA/NCI NIH HHS/United States
- R21 AG042003/AG/NIA NIH HHS/United States
- R41HG007336/HG/NHGRI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

