Single-molecule analysis of DNA replication reveals novel features in the divergent eukaryotes Leishmania and Trypanosoma brucei versus mammalian cells
- PMID: 26976742
- PMCID: PMC4791591
- DOI: 10.1038/srep23142
Single-molecule analysis of DNA replication reveals novel features in the divergent eukaryotes Leishmania and Trypanosoma brucei versus mammalian cells
Abstract
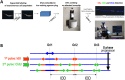
Leishmania and Trypanosoma are unicellular parasites that possess markedly original biological features as compared to other eukaryotes. The Leishmania genome displays a constitutive 'mosaic aneuploidy', whereas in Trypanosoma brucei, the megabase-sized chromosomes are diploid. We accurately analysed DNA replication parameters in three Leishmania species and Trypanosoma brucei as well as mouse embryonic fibroblasts (MEF). Active replication origins were visualized at the single molecule level using DNA molecular combing. More than one active origin was found on most DNA fibres, showing that the chromosomes are replicated from multiple origins. Inter-origin distances (IODs) were measured and found very large in trypanosomatids: the mean IOD was 160 kb in T. brucei and 226 kb in L. mexicana. Moreover, the progression of replication forks was faster than in any other eukaryote analyzed so far (mean velocity 1.9 kb/min in T. brucei and 2.4-2.6 kb/min in Leishmania). The estimated total number of active DNA replication origins in trypanosomatids is ~170. Finally, 14.4% of unidirectional replication forks were observed in T. brucei, in contrast to 1.5-1.7% in Leishmania and 4% in MEF cells. The biological significance of these original features is discussed.
Figures





Comment in
-
Leishmania DNA Replication Timing: A Stochastic Event?Trends Parasitol. 2016 Oct;32(10):755-757. doi: 10.1016/j.pt.2016.05.011. Epub 2016 May 30. Trends Parasitol. 2016. PMID: 27255527
References
-
- Mechali M. DNA replication origins: from sequence specificity to epigenetics. Nat Rev Genet. 2, 640–645 (2001). - PubMed
-
- Lemaitre J. M., Vassetzky Y. & Méchali M. Analysis of Chromatin Assembly, Chromatin Domains, and DNA Replication Using Xenopus Systems. In, Mapping Protein/DNA Interactions by Cross-Linking [Internet]. Paris: Institut national de la santé et de la recherche médicale; 2001. Available from, http://www.ncbi.nlm.nih.gov.gate1.inist.fr/books/NBK7096/ (Accessed: 8th.... - PubMed
-
- Mechali M., Yoshida K., Coulombe P. & Pasero P. Genetic and epigenetic determinants of DNA replication origins, position and activation. Curr Opin Genet Dev. 23, 124–131 (2013). - PubMed
-
- Baldauf S. L. The deep roots of eukaryotes. Science 300, 1703–1706 (2003). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

