Kinetic, Mutational, and Structural Studies of the Venezuelan Equine Encephalitis Virus Nonstructural Protein 2 Cysteine Protease
- PMID: 27030368
- PMCID: PMC5290728
- DOI: 10.1021/acs.biochem.5b00992
Kinetic, Mutational, and Structural Studies of the Venezuelan Equine Encephalitis Virus Nonstructural Protein 2 Cysteine Protease
Abstract
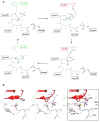
The Venezuelan equine encephalitis virus (VEEV) nonstructural protein 2 (nsP2) cysteine protease (EC 3.4.22.-) is essential for viral replication and is involved in the cytopathic effects (CPE) of the virus. The VEEV nsP2 protease is a member of MEROPS Clan CN and characteristically contains a papain-like protease linked to an S-adenosyl-l-methionine-dependent RNA methyltransferase (SAM MTase) domain. The protease contains an alternative active site motif, (475)NVCWAK(480), which differs from papain's (CGS(25)CWAFS), and the enzyme lacks a transition state-stabilizing residue homologous to Gln-19 in papain. To understand the roles of conserved residues in catalysis, we determined the structure of the free enzyme and the first structure of an inhibitor-bound alphaviral protease. The peptide-like E64d inhibitor was found to bind beneath a β-hairpin at the interface of the SAM MTase and protease domains. His-546 adopted a conformation that differed from that found in the free enzyme; one or both of the conformers may assist in leaving group departure of either the amine or Cys thiolate during the catalytic cycle. Interestingly, E64c (200 μM), the carboxylic acid form of the E64d ester, did not inhibit the nsP2 protease. To identify key residues involved in substrate binding, a number of mutants were analyzed. Mutation of the motif residue, N475A, led to a 24-fold reduction in kcat/Km, and the conformation of this residue did not change after inhibition. N475 forms a hydrogen bond with R662 in the SAM MTase domain, and the R662A and R662K mutations both led to 16-fold decreases in kcat/Km. N475 forms the base of the P1 binding site and likely orients the substrate for nucleophilic attack or plays a role in product release. An Asn homologous to N475 is similarly found in coronaviral papain-like proteases (PLpro) of the Severe Acute Respiratory Syndrome (SARS) virus and Middle East Respiratory Syndrome (MERS) virus. Mutation of another motif residue, K480A, led to a 9-fold decrease in kcat and kcat/Km. K480 likely enhances the nucleophilicity of the Cys. Consistent with our substrate-bound models, the SAM MTase domain K706A mutation increased Km 4.5-fold to 500 μM. Within the β-hairpin, the N545A mutation slightly but not significantly increased kcat and Km. The structures and identified active site residues may facilitate the discovery of protease inhibitors with antiviral activity.
Figures



Similar articles
-
Mutation of Asn-475 in the Venezuelan Equine Encephalitis Virus nsP2 Cysteine Protease Leads to a Self-Inhibited State.Biochemistry. 2017 Nov 28;56(47):6221-6230. doi: 10.1021/acs.biochem.7b00746. Epub 2017 Nov 9. Biochemistry. 2017. PMID: 29064679
-
The crystal structure of the Venezuelan equine encephalitis alphavirus nsP2 protease.Structure. 2006 Sep;14(9):1449-58. doi: 10.1016/j.str.2006.07.010. Structure. 2006. PMID: 16962975
-
Self-inhibited State of Venezuelan Equine Encephalitis Virus (VEEV) nsP2 Cysteine Protease: A Crystallographic and Molecular Dynamics Analysis.J Mol Biol. 2023 Mar 15;435(6):168012. doi: 10.1016/j.jmb.2023.168012. Epub 2023 Feb 13. J Mol Biol. 2023. PMID: 36792007 Free PMC article.
-
Design and Evaluation of Anti-SARS-Coronavirus Agents Based on Molecular Interactions with the Viral Protease.Molecules. 2020 Aug 27;25(17):3920. doi: 10.3390/molecules25173920. Molecules. 2020. PMID: 32867349 Free PMC article. Review.
-
The SARS-coronavirus papain-like protease: structure, function and inhibition by designed antiviral compounds.Antiviral Res. 2015 Mar;115:21-38. doi: 10.1016/j.antiviral.2014.12.015. Epub 2014 Dec 29. Antiviral Res. 2015. PMID: 25554382 Free PMC article. Review.
Cited by
-
Nonstructural Proteins of Alphavirus-Potential Targets for Drug Development.Viruses. 2018 Feb 9;10(2):71. doi: 10.3390/v10020071. Viruses. 2018. PMID: 29425115 Free PMC article. Review.
-
Proteolytic cleavage of host proteins by the Group IV viral proteases of Venezuelan equine encephalitis virus and Zika virus.Antiviral Res. 2019 Apr;164:106-122. doi: 10.1016/j.antiviral.2019.02.001. Epub 2019 Feb 10. Antiviral Res. 2019. PMID: 30742841 Free PMC article.
-
Structural insights into the inhibition of the nsP2 protease from Chikungunya virus by molecular modeling approaches.J Mol Model. 2022 Sep 12;28(10):311. doi: 10.1007/s00894-022-05316-3. J Mol Model. 2022. PMID: 36097090
-
Papain-like cysteine proteinase zone (PCP-zone) and PCP structural catalytic core (PCP-SCC) of enzymes with cysteine proteinase fold.Int J Biol Macromol. 2020 Dec 15;165(Pt A):1438-1446. doi: 10.1016/j.ijbiomac.2020.10.022. Epub 2020 Oct 12. Int J Biol Macromol. 2020. PMID: 33058970 Free PMC article.
-
Specificity Studies of the Venezuelan Equine Encephalitis Virus Non-Structural Protein 2 Protease Using Recombinant Fluorescent Substrates.Int J Mol Sci. 2020 Oct 16;21(20):7686. doi: 10.3390/ijms21207686. Int J Mol Sci. 2020. PMID: 33081394 Free PMC article.
References
-
- Johnson KM, Martin DH. Venezuelan equine encephalitis. Adv Vet Sci Comp Med. 1974;18:79–116. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

