S-nitrosation of proteins relevant to Alzheimer's disease during early stages of neurodegeneration
- PMID: 27035958
- PMCID: PMC4839463
- DOI: 10.1073/pnas.1521318113
S-nitrosation of proteins relevant to Alzheimer's disease during early stages of neurodegeneration
Abstract
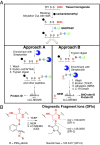
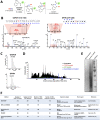
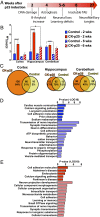
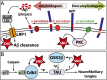
Protein S-nitrosation (SNO-protein), the nitric oxide-mediated posttranslational modification of cysteine thiols, is an important regulatory mechanism of protein function in both physiological and pathological pathways. A key first step toward elucidating the mechanism by which S-nitrosation modulates a protein's function is identification of the targeted cysteine residues. Here, we present a strategy for the simultaneous identification of SNO-cysteine sites and their cognate proteins to profile the brain of the CK-p25-inducible mouse model of Alzheimer's disease-like neurodegeneration. The approach-SNOTRAP (SNO trapping by triaryl phosphine)-is a direct tagging strategy that uses phosphine-based chemical probes, allowing enrichment of SNO-peptides and their identification by liquid chromatography tandem mass spectrometry. SNOTRAP identified 313 endogenous SNO-sites in 251 proteins in the mouse brain, of which 135 SNO-proteins were detected only during neurodegeneration. S-nitrosation in the brain shows regional differences and becomes elevated during early stages of neurodegeneration in the CK-p25 mouse. The SNO-proteome during early neurodegeneration identified increased S-nitrosation of proteins important for synapse function, metabolism, and Alzheimer's disease pathology. In the latter case, proteins related to amyloid precursor protein processing and secretion are S-nitrosated, correlating with increased amyloid formation. Sequence analysis of SNO-cysteine sites identified potential linear motifs that are altered under pathological conditions. Collectively, SNOTRAP is a direct tagging tool for global elucidation of the SNO-proteome, providing functional insights of endogenous SNO proteins in the brain and its dysregulation during neurodegeneration.
Keywords: Alzheimer’s disease; S-nitrosation; neurodegeneration; presenilin pathway; secretase pathway.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
SNOSID, a proteomic method for identification of cysteine S-nitrosylation sites in complex protein mixtures.Proc Natl Acad Sci U S A. 2006 Jan 24;103(4):1012-7. doi: 10.1073/pnas.0508412103. Epub 2006 Jan 17. Proc Natl Acad Sci U S A. 2006. PMID: 16418269 Free PMC article.
-
Site-mapping of in vitro S-nitrosation in cardiac mitochondria: implications for cardioprotection.Mol Cell Proteomics. 2011 Mar;10(3):M110.004721. doi: 10.1074/mcp.M110.004721. Epub 2010 Oct 29. Mol Cell Proteomics. 2011. PMID: 21036925 Free PMC article.
-
An improved sulfur-nitroso-proteome strategy for global profiling of sulfur-nitrosylated proteins and sulfur-nitrosylation sites in mice.J Chromatogr A. 2023 Aug 30;1705:464162. doi: 10.1016/j.chroma.2023.464162. Epub 2023 Jun 15. J Chromatogr A. 2023. PMID: 37336129
-
Protein S-Nitrosation: Biochemistry, Identification, Molecular Mechanisms, and Therapeutic Applications.J Med Chem. 2022 Apr 28;65(8):5902-5925. doi: 10.1021/acs.jmedchem.1c02194. Epub 2022 Apr 12. J Med Chem. 2022. PMID: 35412827 Review.
-
Network analysis of S-nitrosylated synaptic proteins demonstrates unique roles in health and disease.Biochim Biophys Acta Mol Cell Res. 2024 Jun;1871(5):119720. doi: 10.1016/j.bbamcr.2024.119720. Epub 2024 Apr 4. Biochim Biophys Acta Mol Cell Res. 2024. PMID: 38582237 Review.
Cited by
-
The chemical biology of dinitrogen trioxide.Redox Biochem Chem. 2024 Jun;8:100026. doi: 10.1016/j.rbc.2024.100026. Epub 2024 May 9. Redox Biochem Chem. 2024. PMID: 38957295 Free PMC article.
-
S-Glutathionylation and S-Nitrosylation in Mitochondria: Focus on Homeostasis and Neurodegenerative Diseases.Int J Mol Sci. 2022 Dec 13;23(24):15849. doi: 10.3390/ijms232415849. Int J Mol Sci. 2022. PMID: 36555492 Free PMC article. Review.
-
Synthetic PPAR Agonist DTMB Alleviates Alzheimer's Disease Pathology by Inhibition of Chronic Microglial Inflammation in 5xFAD Mice.Neurotherapeutics. 2022 Sep;19(5):1546-1565. doi: 10.1007/s13311-022-01275-y. Epub 2022 Aug 2. Neurotherapeutics. 2022. PMID: 35917087 Free PMC article.
-
Nitric oxide‑mediated S‑Nitrosylation contributes to signaling transduction in human physiological and pathological status (Review).Int J Mol Med. 2025 Oct;56(4):152. doi: 10.3892/ijmm.2025.5593. Epub 2025 Jul 25. Int J Mol Med. 2025. PMID: 40709391 Free PMC article. Review.
-
Metabolic Bypass Rescues Aberrant S-nitrosylation-Induced TCA Cycle Inhibition and Synapse Loss in Alzheimer's Disease Human Neurons.Adv Sci (Weinh). 2024 Mar;11(12):e2306469. doi: 10.1002/advs.202306469. Epub 2024 Jan 18. Adv Sci (Weinh). 2024. PMID: 38235614 Free PMC article.
References
-
- Hess DT, Matsumoto A, Kim SO, Marshall HE, Stamler JS. Protein S-nitrosylation: Purview and parameters. Nat Rev Mol Cell Biol. 2005;6(2):150–166. - PubMed
-
- Martínez-Ruiz A, Cadenas S, Lamas S. Nitric oxide signaling: Classical, less classical, and nonclassical mechanisms. Free Radic Biol Med. 2011;51(1):17–29. - PubMed
-
- Keshive M, Singh S, Wishnok JS, Tannenbaum SR, Deen WM. Kinetics of S-nitrosation of thiols in nitric oxide solutions. Chem Res Toxicol. 1996;9(6):988–993. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

