Smooth Muscle Enriched Long Noncoding RNA (SMILR) Regulates Cell Proliferation
- PMID: 27052414
- PMCID: PMC4872641
- DOI: 10.1161/CIRCULATIONAHA.115.021019
Smooth Muscle Enriched Long Noncoding RNA (SMILR) Regulates Cell Proliferation
Abstract
Background: Phenotypic switching of vascular smooth muscle cells from a contractile to a synthetic state is implicated in diverse vascular pathologies, including atherogenesis, plaque stabilization, and neointimal hyperplasia. However, very little is known about the role of long noncoding RNA (lncRNA) during this process. Here, we investigated a role for lncRNAs in vascular smooth muscle cell biology and pathology.
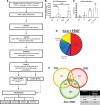
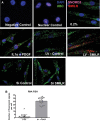
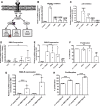
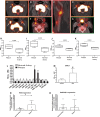
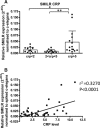
Methods and results: Using RNA sequencing, we identified >300 lncRNAs whose expression was altered in human saphenous vein vascular smooth muscle cells following stimulation with interleukin-1α and platelet-derived growth factor. We focused on a novel lncRNA (Ensembl: RP11-94A24.1), which we termed smooth muscle-induced lncRNA enhances replication (SMILR). Following stimulation, SMILR expression was increased in both the nucleus and cytoplasm, and was detected in conditioned media. Furthermore, knockdown of SMILR markedly reduced cell proliferation. Mechanistically, we noted that expression of genes proximal to SMILR was also altered by interleukin-1α/platelet-derived growth factor treatment, and HAS2 expression was reduced by SMILR knockdown. In human samples, we observed increased expression of SMILR in unstable atherosclerotic plaques and detected increased levels in plasma from patients with high plasma C-reactive protein.
Conclusions: These results identify SMILR as a driver of vascular smooth muscle cell proliferation and suggest that modulation of SMILR may be a novel therapeutic strategy to reduce vascular pathologies.
Keywords: RNA, untranslated; atherosclerosis; cell proliferation; microRNAs; plasma protein, human.
© 2016 The Authors.
Figures



References
-
- Ghouri N, Purves D, McConnachie A, Wilson J, Gill JM, Sattar N. Lower cardiorespiratory fitness contributes to increased insulin resistance and fasting glycaemia in middle-aged South Asian compared with European men living in the UK. Diabetologia. 2013;56:2238–2249. doi: 10.1007/s00125-013-2969-y. - PMC - PubMed
-
- Alexander MR, Owens GK. Epigenetic control of smooth muscle cell differentiation and phenotypic switching in vascular development and disease. Annu Rev Physiol. 2012;74:13–40. doi: 10.1146/annurev-physiol-012110-142315. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

