Nitrite Reductase Activity in Engineered Azurin Variants
- PMID: 27055058
- PMCID: PMC4894363
- DOI: 10.1021/acs.inorgchem.5b03006
Nitrite Reductase Activity in Engineered Azurin Variants
Abstract
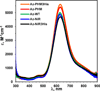
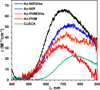
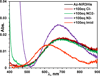
Nitrite reductase (NiR) activity was examined in a series of dicopper P.a. azurin variants in which a surface binding copper site was added through site-directed mutagenesis. Four variants were synthesized with copper binding motifs inspired by the catalytic type 2 copper binding sites found in the native noncoupled dinuclear copper enzymes nitrite reductase and peptidylglycine α-hydroxylating monooxygenase. The four azurin variants, denoted Az-NiR, Az-NiR3His, Az-PHM, and Az-PHM3His, maintained the azurin electron transfer copper center, with the second designed copper site located over 13 Å away and consisting of mutations Asn10His,Gln14Asp,Asn16His-azurin, Asn10His,Gln14His,Asn16His-azurin, Gln8Met,Gln14His,Asn16His-azurin, and Gln8His,Gln14His,Asn16His-azurin, respectively. UV-visible absorption spectroscopy, EPR spectroscopy, and electrochemistry of the sites demonstrate copper binding as well as interaction with small exogenous ligands. The nitrite reduction activity of the variants was determined, including the catalytic Michaelis-Menten parameters. The variants showed activity (0.34-0.59 min(-1)) that was slower than that of native NiRs but comparable to that of other model systems. There were small variations in activity of the four variants that correlated with the number of histidines in the added copper site. Catalysis was found to be reversible, with nitrite produced from NO. Reactions starting with reduced azurin variants demonstrated that electrons from both copper centers were used to reduce nitrite, although steady-state catalysis required the T2 copper center and did not require the T1 center. Finally, experiments separating rates of enzyme reduction from rates of reoxidation by nitrite demonstrated that the reaction with nitrite was rate limiting during catalysis.
Figures


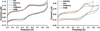






References
-
- Nar H, Messerschmidt A, Huber R, van der Kamp M, Canters GW. J. Mol. Biol. 1991;221:765–772. - PubMed
-
- Suzuki S, Kataoka K, Yamaguchi K. Acc. Chem. Res. 2000;33:728–735. - PubMed
-
- Dodd FE, van Beeumen J, Eady RR, Hasnain SS. J. Mol. Biol. 1998;282:369–382. - PubMed
-
- Adman ET, Murphy MEP. Copper Nitrite Reductase. In: Messerschmid A, Huber R, Poulos T, Wieghardt K, editors. Handbook of Metalloproteins. Vol. 2. Chichester, UK: Wiley; 2001. pp. 1381–1390.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

