RNA Polymerase II Regulates Topoisomerase 1 Activity to Favor Efficient Transcription
- PMID: 27058666
- PMCID: PMC4826470
- DOI: 10.1016/j.cell.2016.02.036
RNA Polymerase II Regulates Topoisomerase 1 Activity to Favor Efficient Transcription
Abstract
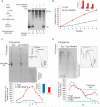
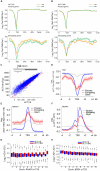
We report a mechanism through which the transcription machinery directly controls topoisomerase 1 (TOP1) activity to adjust DNA topology throughout the transcription cycle. By comparing TOP1 occupancy using chromatin immunoprecipitation sequencing (ChIP-seq) versus TOP1 activity using topoisomerase 1 sequencing (TOP1-seq), a method reported here to map catalytically engaged TOP1, TOP1 bound at promoters was discovered to become fully active only after pause-release. This transition coupled the phosphorylation of the carboxyl-terminal-domain (CTD) of RNA polymerase II (RNAPII) with stimulation of TOP1 above its basal rate, enhancing its processivity. TOP1 stimulation is strongly dependent on the kinase activity of BRD4, a protein that phosphorylates Ser2-CTD and regulates RNAPII pause-release. Thus the coordinated action of BRD4 and TOP1 overcame the torsional stress opposing transcription as RNAPII commenced elongation but preserved negative supercoiling that assists promoter melting at start sites. This nexus between transcription and DNA topology promises to elicit new strategies to intercept pathological gene expression.
Copyright © 2016 Elsevier Inc. All rights reserved.
Figures





References
-
- Amente S, Gargano B, Napolitano G, Lania L, Majello B. Camptothecin releases P-TEFb from the inactive 7SK snRNP complex. Cell Cycle. 2009;8:1249–1255. - PubMed
-
- Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z, Wei G, Chepelev I, Zhao K. High-resolution profiling of histone methylations in the human genome. Cell. 2007;129:823–837. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

