Consequences of Cas9 cleavage in the chromosome of Escherichia coli
- PMID: 27060147
- PMCID: PMC4872102
- DOI: 10.1093/nar/gkw223
Consequences of Cas9 cleavage in the chromosome of Escherichia coli
Abstract
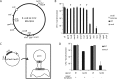
The RNA-guided Cas9 nuclease from CRISPR-Cas systems has emerged as a powerful biotechnological tool. The specificity of Cas9 can be reprogrammed to cleave desired sequences in a cell's chromosome simply by changing the sequence of a small guide RNA. Unlike in most eukaryotes, Cas9 cleavage in the chromosome of bacteria has been reported to kill the cell. However, the mechanism of cell death remains to be investigated. Bacteria mainly rely on homologous recombination (HR) with sister chromosomes to repair double strand breaks. Here, we show that the simultaneous cleavage of all copies of the Escherichia coli chromosome at the same position cannot be repaired, leading to cell death. However, inefficient cleavage can be tolerated through continuous repair by the HR pathway. In order to kill cells reliably, HR can be blocked using the Mu phage Gam protein. Finally, the introduction of the non-homologous end joining (NHEJ) pathway from Mycobacterium tuberculosis was not able to rescue the cells from Cas9-mediated killing, but did introduce small deletions at a low frequency. This work provides a better understanding of the consequences of Cas9 cleavage in bacterial chromosomes which will be instrumental in the development of future CRISPR tools.
© The Author(s) 2016. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
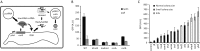



Similar articles
-
CRISPR-Cas: New Tools for Genetic Manipulations from Bacterial Immunity Systems.Annu Rev Microbiol. 2015;69:209-28. doi: 10.1146/annurev-micro-091014-104441. Epub 2015 Jul 22. Annu Rev Microbiol. 2015. PMID: 26209264 Review.
-
An efficient system for deletion of large DNA fragments in Escherichia coli via introduction of both Cas9 and the non-homologous end joining system from Mycobacterium smegmatis.Biochem Biophys Res Commun. 2017 Apr 15;485(4):768-774. doi: 10.1016/j.bbrc.2017.02.129. Epub 2017 Feb 28. Biochem Biophys Res Commun. 2017. PMID: 28257845
-
CRISPR-Cas9-assisted native end-joining editing offers a simple strategy for efficient genetic engineering in Escherichia coli.Appl Microbiol Biotechnol. 2019 Oct;103(20):8497-8509. doi: 10.1007/s00253-019-10104-w. Epub 2019 Sep 9. Appl Microbiol Biotechnol. 2019. PMID: 31501938
-
A CRISPR-Cas9 Assisted Non-Homologous End-Joining Strategy for One-step Engineering of Bacterial Genome.Sci Rep. 2016 Nov 24;6:37895. doi: 10.1038/srep37895. Sci Rep. 2016. PMID: 27883076 Free PMC article.
-
Exploring the potential of genome editing CRISPR-Cas9 technology.Gene. 2017 Jan 30;599:1-18. doi: 10.1016/j.gene.2016.11.008. Epub 2016 Nov 9. Gene. 2017. PMID: 27836667 Review.
Cited by
-
Intein-based thermoregulated meganucleases for containment of genetic material.Nucleic Acids Res. 2024 Feb 28;52(4):2066-2077. doi: 10.1093/nar/gkad1247. Nucleic Acids Res. 2024. PMID: 38180814 Free PMC article.
-
Recombineering and MAGE.Nat Rev Methods Primers. 2021;1:7. doi: 10.1038/s43586-020-00006-x. Epub 2021 Jan 14. Nat Rev Methods Primers. 2021. PMID: 35540496 Free PMC article.
-
A bacterial gene-drive system efficiently edits and inactivates a high copy number antibiotic resistance locus.Nat Commun. 2019 Dec 16;10(1):5726. doi: 10.1038/s41467-019-13649-6. Nat Commun. 2019. PMID: 31844051 Free PMC article.
-
Genome-wide CRISPR-dCas9 screens in E. coli identify essential genes and phage host factors.PLoS Genet. 2018 Nov 7;14(11):e1007749. doi: 10.1371/journal.pgen.1007749. eCollection 2018 Nov. PLoS Genet. 2018. PMID: 30403660 Free PMC article.
-
Barriers to genome editing with CRISPR in bacteria.J Ind Microbiol Biotechnol. 2019 Oct;46(9-10):1327-1341. doi: 10.1007/s10295-019-02195-1. Epub 2019 Jun 5. J Ind Microbiol Biotechnol. 2019. PMID: 31165970 Free PMC article. Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

