The genetic diversity of triticale genotypes involved in Polish breeding programs
- PMID: 27066368
- PMCID: PMC4801839
- DOI: 10.1186/s40064-016-1997-8
The genetic diversity of triticale genotypes involved in Polish breeding programs
Abstract
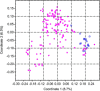
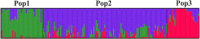
Genetic diversity analysis of triticale populations is useful for breeding programs, as it helps to select appropriate genetic material for classifying the parental lines, heterotic groups and predicting hybrid performance. In our study 232 breeding forms were analyzed using diversity arrays technology markers. Principal coordinate analysis followed by model-based Bayesian analysis of population structure revealed the presence of weak data structuring with three groups of data. In the first group, 17 spring and 17 winter forms were clustered. The second and the third groups were represented by 101 and 26 winter forms, respectively. Polymorphic information content values, as well as Shannon's Information Index, were higher for the first (0.319) and second (0.309) than for third (0.234) group. AMOVA analysis demonstrated a higher level of within variation (86 %) than among populations (14 %). This study provides the basic information on the presence of structure within a genetic pool of triticale breeding forms.
Keywords: DArT markers; Genetic diversity; Triticale.
Figures
Similar articles
-
Genetic diversity in European winter triticale determined with SSR markers and coancestry coefficient.Theor Appl Genet. 2004 May;108(7):1385-91. doi: 10.1007/s00122-003-1552-1. Epub 2004 Feb 4. Theor Appl Genet. 2004. PMID: 14760487
-
Analysis of the Genetic Diversity and Population Structure of Austrian and Belgian Wheat Germplasm within a Regional Context Based on DArT Markers.Genes (Basel). 2018 Jan 22;9(1):47. doi: 10.3390/genes9010047. Genes (Basel). 2018. PMID: 29361778 Free PMC article.
-
Assessment of genetic diversity among sunflower genotypes using microsatellite markers.Mol Biol Res Commun. 2018 Sep;7(3):143-152. doi: 10.22099/mbrc.2018.30434.1340. Mol Biol Res Commun. 2018. PMID: 30426032 Free PMC article.
-
AFLP-Based Analysis of Genetic Diversity, Population Structure, and Relationships with Agronomic Traits in Rice Germplasm from North Region of Iran and World Core Germplasm Set.Biochem Genet. 2016 Apr;54(2):177-93. doi: 10.1007/s10528-016-9711-7. Epub 2016 Jan 13. Biochem Genet. 2016. PMID: 26762294
-
Genetic Diversity and Population Structure Analysis of DalbergiaOdorifera Germplasm and Development of a Core Collection Using Microsatellite Markers.Genes (Basel). 2019 Apr 6;10(4):281. doi: 10.3390/genes10040281. Genes (Basel). 2019. PMID: 30959931 Free PMC article.
Cited by
-
Effect of Chromosomal Localization of NGS-Based Markers on Their Applicability for Analyzing Genetic Variation and Population Structure of Hexaploid Triticale.Int J Mol Sci. 2024 Sep 3;25(17):9568. doi: 10.3390/ijms25179568. Int J Mol Sci. 2024. PMID: 39273515 Free PMC article.
-
Pyramiding wheat pre-harvest sprouting resistance genes in triticale breeding.Mol Breed. 2022 Sep 22;42(10):60. doi: 10.1007/s11032-022-01327-3. eCollection 2022 Oct. Mol Breed. 2022. PMID: 37309488 Free PMC article.
-
Genetic mapping of male sterility and pollen fertility QTLs in triticale with sterilizing Triticum timopheevii cytoplasm.J Appl Genet. 2021 Feb;62(1):59-71. doi: 10.1007/s13353-020-00595-z. Epub 2020 Nov 23. J Appl Genet. 2021. PMID: 33230679 Free PMC article.
-
Assessing the genetic diversity and characterizing genomic regions conferring Tan Spot resistance in cultivated rye.PLoS One. 2019 Mar 28;14(3):e0214519. doi: 10.1371/journal.pone.0214519. eCollection 2019. PLoS One. 2019. PMID: 30921415 Free PMC article.
-
Genome-wide association study for in vitro digestibility and related traits in triticale forage.BMC Plant Biol. 2024 Mar 27;24(1):223. doi: 10.1186/s12870-024-04927-7. BMC Plant Biol. 2024. PMID: 38539072 Free PMC article.
References
-
- Alheit KV, Reif JC, Maurer HP, Hahn V, Weissmann EA, Miedaner T, Würschum T. Detection of segregation distortion loci in triticale (x Triticosecale Wittmack) based on a high-density DArT marker consensus genetic linkage map. BMC Genom. 2011;12:380. doi: 10.1186/1471-2164-12-380. - DOI - PMC - PubMed
-
- Ammar K, Mergoum M, Rajaram S. The history and evolution of triticale. In: Mergoum M, Gómez-Macpherson H, editors. Triticale improvement and production. Rome: Food and Agriculture Organization of the United Nations; 2004. pp. 1–10.
LinkOut - more resources
Full Text Sources
Other Literature Sources

