Overlapping Community Detection based on Network Decomposition
- PMID: 27066904
- PMCID: PMC4828636
- DOI: 10.1038/srep24115
Overlapping Community Detection based on Network Decomposition
Abstract
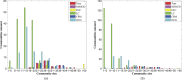
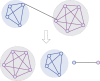
Community detection in complex network has become a vital step to understand the structure and dynamics of networks in various fields. However, traditional node clustering and relatively new proposed link clustering methods have inherent drawbacks to discover overlapping communities. Node clustering is inadequate to capture the pervasive overlaps, while link clustering is often criticized due to the high computational cost and ambiguous definition of communities. So, overlapping community detection is still a formidable challenge. In this work, we propose a new overlapping community detection algorithm based on network decomposition, called NDOCD. Specifically, NDOCD iteratively splits the network by removing all links in derived link communities, which are identified by utilizing node clustering technique. The network decomposition contributes to reducing the computation time and noise link elimination conduces to improving the quality of obtained communities. Besides, we employ node clustering technique rather than link similarity measure to discover link communities, thus NDOCD avoids an ambiguous definition of community and becomes less time-consuming. We test our approach on both synthetic and real-world networks. Results demonstrate the superior performance of our approach both in computation time and accuracy compared to state-of-the-art algorithms.
Figures






References
-
- Newman M. E. J. Communities, modules and large-scale structure in networks. Nat. Phys. 8, 25–31 (2012).
-
- Fortunato S. Community detection in graphs. Phys. Rep. 486, 75–174 (2010).
-
- Xie J., Kelley S. & Szymanski B. K. Overlapping community detection in networks: the state of the art and comparative study. ACM Comput. Surv. 45, 43, 10.1145/2501654.2501657 (2013). - DOI
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

