Nested Russian Doll-Like Genetic Mobility Drives Rapid Dissemination of the Carbapenem Resistance Gene blaKPC
- PMID: 27067320
- PMCID: PMC4879409
- DOI: 10.1128/AAC.00464-16
Nested Russian Doll-Like Genetic Mobility Drives Rapid Dissemination of the Carbapenem Resistance Gene blaKPC
Abstract
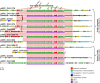
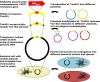
The recent widespread emergence of carbapenem resistance in Enterobacteriaceae is a major public health concern, as carbapenems are a therapy of last resort against this family of common bacterial pathogens. Resistance genes can mobilize via various mechanisms, including conjugation and transposition; however, the importance of this mobility in short-term evolution, such as within nosocomial outbreaks, is unknown. Using a combination of short- and long-read whole-genome sequencing of 281 blaKPC-positive Enterobacteriaceae isolates from a single hospital over 5 years, we demonstrate rapid dissemination of this carbapenem resistance gene to multiple species, strains, and plasmids. Mobility of blaKPC occurs at multiple nested genetic levels, with transmission of blaKPC strains between individuals, frequent transfer of blaKPC plasmids between strains/species, and frequent transposition of blaKPC transposon Tn4401 between plasmids. We also identify a common insertion site for Tn4401 within various Tn2-like elements, suggesting that homologous recombination between Tn2-like elements has enhanced the spread of Tn4401 between different plasmid vectors. Furthermore, while short-read sequencing has known limitations for plasmid assembly, various studies have attempted to overcome this by the use of reference-based methods. We also demonstrate that, as a consequence of the genetic mobility observed in this study, plasmid structures can be extremely dynamic, and therefore these reference-based methods, as well as traditional partial typing methods, can produce very misleading conclusions. Overall, our findings demonstrate that nonclonal resistance gene dissemination can be extremely rapid, presenting significant challenges for public health surveillance and achieving effective control of antibiotic resistance.
Copyright © 2016 Sheppard et al.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

