The tandem duplicator phenotype as a distinct genomic configuration in cancer
- PMID: 27071093
- PMCID: PMC4855596
- DOI: 10.1073/pnas.1520010113
The tandem duplicator phenotype as a distinct genomic configuration in cancer
Abstract
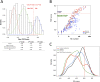
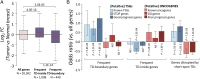
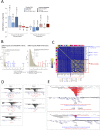
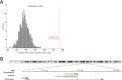
Next-generation sequencing studies have revealed genome-wide structural variation patterns in cancer, such as chromothripsis and chromoplexy, that do not engage a single discernable driver mutation, and whose clinical relevance is unclear. We devised a robust genomic metric able to identify cancers with a chromotype called tandem duplicator phenotype (TDP) characterized by frequent and distributed tandem duplications (TDs). Enriched only in triple-negative breast cancer (TNBC) and in ovarian, endometrial, and liver cancers, TDP tumors conjointly exhibit tumor protein p53 (TP53) mutations, disruption of breast cancer 1 (BRCA1), and increased expression of DNA replication genes pointing at rereplication in a defective checkpoint environment as a plausible causal mechanism. The resultant TDs in TDP augment global oncogene expression and disrupt tumor suppressor genes. Importantly, the TDP strongly correlates with cisplatin sensitivity in both TNBC cell lines and primary patient-derived xenografts. We conclude that the TDP is a common cancer chromotype that coordinately alters oncogene/tumor suppressor expression with potential as a marker for chemotherapeutic response.
Keywords: BRCA1; TP53; cisplatin; tandem duplications; triple-negative breast cancer.
Conflict of interest statement
The authors declare no conflict of interest.
Figures







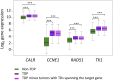


Comment in
-
Tandem duplications contribute to not one but two distinct phenotypes.Proc Natl Acad Sci U S A. 2016 Sep 6;113(36):E5257-8. doi: 10.1073/pnas.1610228113. Epub 2016 Aug 19. Proc Natl Acad Sci U S A. 2016. PMID: 27543336 Free PMC article. No abstract available.
References
-
- Hanahan D, Weinberg RA. Hallmarks of cancer: The next generation. Cell. 2011;144(5):646–674. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

