BRONCO: Biomedical entity Relation ONcology COrpus for extracting gene-variant-disease-drug relations
- PMID: 27074804
- PMCID: PMC4830473
- DOI: 10.1093/database/baw043
BRONCO: Biomedical entity Relation ONcology COrpus for extracting gene-variant-disease-drug relations
Abstract
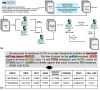
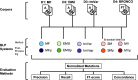
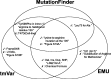
Comprehensive knowledge of genomic variants in a biological context is key for precision medicine. As next-generation sequencing technologies improve, the amount of literature containing genomic variant data, such as new functions or related phenotypes, rapidly increases. Because numerous articles are published every day, it is almost impossible to manually curate all the variant information from the literature. Many researchers focus on creating an improved automated biomedical natural language processing (BioNLP) method that extracts useful variants and their functional information from the literature. However, there is no gold-standard data set that contains texts annotated with variants and their related functions. To overcome these limitations, we introduce a Biomedical entity Relation ONcology COrpus (BRONCO) that contains more than 400 variants and their relations with genes, diseases, drugs and cell lines in the context of cancer and anti-tumor drug screening research. The variants and their relations were manually extracted from 108 full-text articles. BRONCO can be utilized to evaluate and train new methods used for extracting biomedical entity relations from full-text publications, and thus be a valuable resource to the biomedical text mining research community. Using BRONCO, we quantitatively and qualitatively evaluated the performance of three state-of-the-art BioNLP methods. We also identified their shortcomings, and suggested remedies for each method. We implemented post-processing modules for the three BioNLP methods, which improved their performance.Database URL:http://infos.korea.ac.kr/bronco.
© The Author(s) 2016. Published by Oxford University Press.
Figures

References
-
- Kumar P, Henikoff S, Ng PC. (2009) Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 4(7):1073–1082. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

