Relative Quantification of Sites of Peptide and Protein Modification Using Size Exclusion Chromatography Coupled with Electron Transfer Dissociation
- PMID: 27075875
- PMCID: PMC4945384
- DOI: 10.1007/s13361-016-1403-3
Relative Quantification of Sites of Peptide and Protein Modification Using Size Exclusion Chromatography Coupled with Electron Transfer Dissociation
Abstract
One difficult problem in the analysis of peptide modifications is quantifying isomeric modifications that differ by the position of the amino acid modified. HPLC separation using C18 reverse phase chromatography coupled with electron transfer dissociation (ETD) in tandem mass spectrometry has recently been shown to be able to relatively quantify how much of a given modification occurs at each amino acid position for isomeric mixtures; however, the resolution of reverse phase chromatography greatly complicates quantification of isomeric modifications by ETD because of the chromatographic separation of peptides with identical modifications at different sequence positions. Using peptide oxidation as a model system, we investigated the use of size exclusion chromatography coupled with ETD fragmentation to separate peptide sequences. This approach allows for the benefits of chromatographic separation of peptide sequences while ensuring co-elution of modification isomers for accurate relative quantification of modifications using standard data-dependent acquisitions. Using this method, the relative amount of modification at each amino acid can be accurately measured from single ETD MS/MS spectra in a standard data-dependent acquisition experiment. Graphical Abstract ᅟ.
Keywords: Electron transfer dissociation; Fast photochemical oxidation of proteins; Hydroxyl radical protein footprinting; Peptide modification; Size exclusion chromatography.
Figures

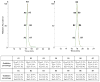

References
-
- Garcia BA, Shabanowitz J, Hunt DF. Analysis of protein phosphorylation by mass spectrometry. Methods. 2005;35(3):256–64. - PubMed
-
- Mann M, Ong SE, Gronborg M, Steen H, Jensen ON, Pandey A. Analysis of protein phosphorylation using mass spectrometry: deciphering the phosphoproteome. Trends Biotechnol. 2002;20(6):261–8. - PubMed
-
- Nagel AK, Schilling M, Comte-Walters S, Berkaw MN, Ball LE. Identification of O-linked N-acetylglucosamine (O-GlcNAc)-modified osteoblast proteins by electron transfer dissociation tandem mass spectrometry reveals proteins critical for bone formation. Mol Cell Proteomics. 2013;12(4):945–55. - PMC - PubMed
-
- Greis KD, Hayes BK, Comer FI, Kirk M, Barnes S, Lowary TL, Hart GW. Selective detection and site-analysis of O-GlcNAc-modified glycopeptides by beta-elimination and tandem electrospray mass spectrometry. Anal Biochem. 1996;234(1):38–49. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

