How Many Parameters Does It Take to Describe Disease Tolerance?
- PMID: 27088212
- PMCID: PMC4835111
- DOI: 10.1371/journal.pbio.1002435
How Many Parameters Does It Take to Describe Disease Tolerance?
Erratum in
-
Correction: How Many Parameters Does It Take to Describe Disease Tolerance?PLoS Biol. 2016 Jun 6;14(6):e1002485. doi: 10.1371/journal.pbio.1002485. eCollection 2016 Jun. PLoS Biol. 2016. PMID: 27270605 Free PMC article.
Abstract
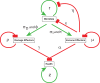
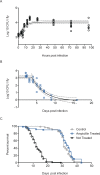
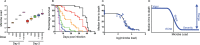
The study of infectious disease has been aided by model organisms, which have helped to elucidate molecular mechanisms and contributed to the development of new treatments; however, the lack of a conceptual framework for unifying findings across models, combined with host variability, has impeded progress and translation. Here, we fill this gap with a simple graphical and mathematical framework to study disease tolerance, the dose response curve relating health to microbe load; this approach helped uncover parameters that were previously overlooked. Using a model experimental system in which we challenged Drosophila melanogaster with the pathogen Listeria monocytogenes, we tested this framework, finding that microbe growth, the immune response, and disease tolerance were all well represented by sigmoid models. As we altered the system by varying host or pathogen genetics, disease tolerance varied, as we would expect if it was indeed governed by parameters controlling the sensitivity of the system (the number of bacteria required to trigger a response) and maximal effect size according to a logistic equation. Though either the pathogen or host immune response or both together could theoretically be the proximal cause of pathology that killed the flies, we found that the pathogen, but not the immune response, drove damage in this model. With this new understanding of the circuitry controlling disease tolerance, we can now propose better ways of choosing, combining, and developing treatments.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Raberg l, Sim D, Read AF. Disentangling genetic variation for resistance and tolerance to infectious diseases in animals. Science. 2007;318:812–4. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

