Global proteome profiling of dental cementum under experimentally-induced apposition
- PMID: 27095596
- PMCID: PMC4908826
- DOI: 10.1016/j.jprot.2016.03.036
Global proteome profiling of dental cementum under experimentally-induced apposition
Abstract
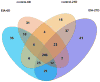
Dental cementum (DC) covers the tooth root and has important functions in tooth attachment and position. DC can be lost to disease, and regeneration is currently unpredictable due to limited understanding of DC formation. This study used a model of experimentally-induced apposition (EIA) in mice to identify proteins associated with new DC formation. Mandibular first molars were induced to super-erupt for 6 and 21days after extracting opposing maxillary molars. Decalcified and formalin-fixed paraffin-embedded mandible sections were prepared for laser capture microdissection. Microdissected protein extracts were analyzed by liquid chromatography coupled to tandem mass spectrometry (LC-MS/MS), and the data submitted to repeated measure ANOVA test (RM-ANOVA, alpha=5%). A total of 519 proteins were identified, with 97 (18.6%) proteins found exclusively in EIA sites and 50 (9.6%) proteins exclusively expressed in control sites. Fifty six (10.7%) proteins were differentially regulated by RM-ANOVA (p<0.05), with 24 regulated by the exclusive effect of EIA (12 proteins) or the interaction between EIA and time (12 proteins), including serpin 1a, procollagen C-endopeptidase enhancer, tenascin X (TNX), and asporin (ASPN). In conclusion, proteomic analysis demonstrated significantly altered protein profile in DC under EIA, providing new insights on DC biology and potential candidates for tissue engineering applications.
Significance: Dental cementum (DC) is a mineralized tissue that covers the tooth root surface and has important functions in tooth attachment and position. DC and other periodontal tissues can be lost to disease, and regeneration is currently unpredictable due to lack of understanding of DC formation. This study used a model of experimentally-induced apposition (EIA) in mice to promote new cementum formation, followed by laser capture microdissection (LCM) and liquid chromatography coupled to tandem mass spectrometry (LC-MS/MS) proteomic analysis. This approach identified proteins associated with new cementum formation that may be targets for promoting cementum regeneration.
Keywords: Dental cementum; Periodontal ligament; Proteomic analysis; Tissue apposition.
Copyright © 2016 Elsevier B.V. All rights reserved.
Conflict of interest statement
The authors of manuscript entitled
The authors report no conflict of interest.
Figures








Similar articles
-
Proteomic analysis of human dental cementum and alveolar bone.J Proteomics. 2013 Oct 8;91:544-55. doi: 10.1016/j.jprot.2013.08.016. Epub 2013 Sep 2. J Proteomics. 2013. PMID: 24007660 Free PMC article.
-
Microproteome of dentoalveolar tissues.Bone. 2017 Aug;101:219-229. doi: 10.1016/j.bone.2017.05.014. Epub 2017 May 17. Bone. 2017. PMID: 28527949
-
Cementocyte alterations associated with experimentally induced cellular cementum apposition in Hyp mice.J Periodontol. 2021 Nov;92(11):116-127. doi: 10.1002/JPER.21-0119. Epub 2021 Jun 9. J Periodontol. 2021. PMID: 34003518 Free PMC article.
-
Cementogenesis reviewed: a comparison between human premolars and rodent molars.Anat Rec. 1996 Jun;245(2):267-92. doi: 10.1002/(SICI)1097-0185(199606)245:2<267::AID-AR12>3.0.CO;2-N. Anat Rec. 1996. PMID: 8769668 Review.
-
Cementum proteins: role in cementogenesis, biomineralization, periodontium formation and regeneration.Periodontol 2000. 2015 Feb;67(1):211-33. doi: 10.1111/prd.12062. Periodontol 2000. 2015. PMID: 25494602 Review.
Cited by
-
Osteopontin regulates dentin and alveolar bone development and mineralization.Bone. 2018 Feb;107:196-207. doi: 10.1016/j.bone.2017.12.004. Epub 2017 Dec 5. Bone. 2018. PMID: 29313816 Free PMC article.
-
Short histological kaleidoscope - recent findings in histology. Part I.Rom J Morphol Embryol. 2022 Jan-Mar;63(1):7-29. doi: 10.47162/RJME.63.1.01. Rom J Morphol Embryol. 2022. PMID: 36074664 Free PMC article. Review.
-
Obesity Modifies the Proteomic Profile of the Periodontal Ligament.Int J Mol Sci. 2023 Jan 5;24(2):1003. doi: 10.3390/ijms24021003. Int J Mol Sci. 2023. PMID: 36674516 Free PMC article.
-
Disparate effects of sclerostin deletion on alveolar bone and cellular cementum in mice.J Periodontol. 2025 Jan;96(1):82-96. doi: 10.1002/JPER.24-0025. Epub 2024 Jul 16. J Periodontol. 2025. PMID: 39012429 Free PMC article.
-
Development of immortalized Hertwig's epithelial root sheath cell lines for cementum and dentin regeneration.Stem Cell Res Ther. 2019 Jan 3;10(1):3. doi: 10.1186/s13287-018-1106-8. Stem Cell Res Ther. 2019. PMID: 30606270 Free PMC article.
References
-
- Foster BL, Somerman MJ. Cementum. In: McCauley LK, Somerman MJ, editors. Mineralized Tissues in Oral and Craniofacial Science: Biological Principles and Clinical Correlates. 1. Ames, IA: Wiley-Blackwell; 2012. pp. 169–92.
-
- Foster BL, Nociti FH, Jr, Somerman MJ. Tooth Root Formation. In: Huang GTJ, Thesleff I, editors. Stem Cells, Craniofacial Development and Regeneration. 1. Wiley-Blackwell; 2013. pp. 153–77.
-
- Bosshardt D. Are cementoblasts a subpopulation of osteoblasts or a unique phenotype? J Dent Res. 2005;84:390–406. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

